+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1g13 | ||||||
|---|---|---|---|---|---|---|---|
| Title | HUMAN GM2 ACTIVATOR STRUCTURE | ||||||
 Components Components | GANGLIOSIDE M2 ACTIVATOR PROTEIN | ||||||
 Keywords Keywords | Ligand Binding Protein / beta cup | ||||||
| Function / homology |  Function and homology information Function and homology informationsphingolipid activator protein activity / beta-N-acetylgalactosaminidase activity / glycosphingolipid catabolic process / lipid transporter activity / Glycosphingolipid catabolism / lipid storage / ganglioside catabolic process / oligosaccharide catabolic process / neuromuscular process controlling balance / phospholipase activator activity ...sphingolipid activator protein activity / beta-N-acetylgalactosaminidase activity / glycosphingolipid catabolic process / lipid transporter activity / Glycosphingolipid catabolism / lipid storage / ganglioside catabolic process / oligosaccharide catabolic process / neuromuscular process controlling balance / phospholipase activator activity / lipid transport / lysosomal lumen / cytoplasmic side of plasma membrane / azurophil granule lumen / basolateral plasma membrane / learning or memory / apical plasma membrane / intracellular membrane-bounded organelle / Neutrophil degranulation / extracellular exosome / extracellular region / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MAD / Resolution: 2 Å MAD / Resolution: 2 Å | ||||||
 Authors Authors | Wright, C.S. / Li, S.C. / Rastinejad, F. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2000 Journal: J.Mol.Biol. / Year: 2000Title: Crystal structure of human GM2-activator protein with a novel beta-cup topology. Authors: Wright, C.S. / Li, S.C. / Rastinejad, F. #1:  Journal: Acta Crystallogr.,Sect.D / Year: 1997 Journal: Acta Crystallogr.,Sect.D / Year: 1997Title: Crystallization and preliminary X-ray characterization of GM2-activator protein Authors: Wright, C.S. / Li, S.-C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1g13.cif.gz 1g13.cif.gz | 112.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1g13.ent.gz pdb1g13.ent.gz | 87.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1g13.json.gz 1g13.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1g13_validation.pdf.gz 1g13_validation.pdf.gz | 451.3 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1g13_full_validation.pdf.gz 1g13_full_validation.pdf.gz | 456.2 KB | Display | |
| Data in XML |  1g13_validation.xml.gz 1g13_validation.xml.gz | 27.9 KB | Display | |
| Data in CIF |  1g13_validation.cif.gz 1g13_validation.cif.gz | 37.7 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/g1/1g13 https://data.pdbj.org/pub/pdb/validation_reports/g1/1g13 ftp://data.pdbj.org/pub/pdb/validation_reports/g1/1g13 ftp://data.pdbj.org/pub/pdb/validation_reports/g1/1g13 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 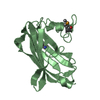
| ||||||||
| 3 | 
| ||||||||
| 4 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 17698.096 Da / Num. of mol.: 3 / Fragment: RESIDUES 39-200 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Organ: KIDNEY / Production host: Homo sapiens (human) / Organ: KIDNEY / Production host:  #2: Chemical | ChemComp-EPE / | #3: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.94 Å3/Da / Density % sol: 58 % | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 7.5 Details: PEG 4000, propanol, sodium Hepes, pH 7.5, VAPOR DIFFUSION, HANGING DROP, temperature 298K | |||||||||||||||||||||||||
| Crystal grow | *PLUS | |||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 150 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  CHESS CHESS  / Beamline: F2 / Wavelength: 0.979 / Beamline: F2 / Wavelength: 0.979 |
| Detector | Type: ccd / Detector: CCD / Date: May 28, 1998 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.979 Å / Relative weight: 1 |
| Reflection | Resolution: 2→20 Å / Num. all: 44590 / Num. obs: 44581 / % possible obs: 99.8 % / Observed criterion σ(F): 1 / Observed criterion σ(I): 1 / Redundancy: 6 % / Biso Wilson estimate: 33.3 Å2 / Rmerge(I) obs: 0.049 / Net I/σ(I): 45.7 |
| Reflection shell | Resolution: 2→2.07 Å / Redundancy: 10 % / Rmerge(I) obs: 0.33 / Num. unique all: 4326 / % possible all: 99 |
| Reflection | *PLUS Num. measured all: 84502 |
| Reflection shell | *PLUS % possible obs: 99 % |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MAD / Resolution: 2→6 Å / σ(F): 3 / σ(I): 3 / Stereochemistry target values: Engh & Huber / Details: torsion angle refinement MAD / Resolution: 2→6 Å / σ(F): 3 / σ(I): 3 / Stereochemistry target values: Engh & Huber / Details: torsion angle refinement
| |||||||||||||||||||||||||||
| Solvent computation | Solvent model: Least Squares / Bsol: 136.55 Å2 / ksol: 0.7133 e/Å3 | |||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 34 Å2 | |||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.24 Å / Luzzati sigma a obs: 0.09 Å | |||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→6 Å
| |||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2→6 Å / Total num. of bins used: 10
| |||||||||||||||||||||||||||
| Software | *PLUS Name: CNS / Version: 0.9 / Classification: refinement | |||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller













 PDBj
PDBj



