+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1esk | ||||||
|---|---|---|---|---|---|---|---|
| Title | SOLUTION STRUCTURE OF NCP7 FROM HIV-1 | ||||||
 Components Components | GAG POLYPROTEIN | ||||||
 Keywords Keywords | VIRAL PROTEIN / (12-53)NCp7 / HIV-1 / PROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology informationintegrase activity / Integration of viral DNA into host genomic DNA / Autointegration results in viral DNA circles / Minus-strand DNA synthesis / Plus-strand DNA synthesis / Uncoating of the HIV Virion / 2-LTR circle formation / Vpr-mediated nuclear import of PICs / Early Phase of HIV Life Cycle / Integration of provirus ...integrase activity / Integration of viral DNA into host genomic DNA / Autointegration results in viral DNA circles / Minus-strand DNA synthesis / Plus-strand DNA synthesis / Uncoating of the HIV Virion / 2-LTR circle formation / Vpr-mediated nuclear import of PICs / Early Phase of HIV Life Cycle / Integration of provirus / APOBEC3G mediated resistance to HIV-1 infection / Binding and entry of HIV virion / viral life cycle / HIV-1 retropepsin / symbiont-mediated activation of host apoptosis / retroviral ribonuclease H / exoribonuclease H / exoribonuclease H activity / Assembly Of The HIV Virion / Budding and maturation of HIV virion / protein processing / viral genome integration into host DNA / establishment of integrated proviral latency / RNA-directed DNA polymerase / RNA stem-loop binding / viral penetration into host nucleus / host multivesicular body / RNA-directed DNA polymerase activity / RNA-DNA hybrid ribonuclease activity / Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / peptidase activity / host cell / viral nucleocapsid / DNA recombination / DNA-directed DNA polymerase / aspartic-type endopeptidase activity / Hydrolases; Acting on ester bonds / DNA-directed DNA polymerase activity / symbiont-mediated suppression of host gene expression / viral translational frameshifting / symbiont entry into host cell / lipid binding / host cell plasma membrane / host cell nucleus / virion membrane / structural molecule activity / DNA binding / zinc ion binding / identical protein binding Similarity search - Function | ||||||
| Method | SOLUTION NMR / simulated annealing | ||||||
 Authors Authors | Morellet, N. / Demene, H. / Teilleux, V. / Huynh-Dinh, T. / de Rocquigny, H. / Fournie-Zaluski, M.-C. / Roques, B.P. | ||||||
 Citation Citation |  Journal: To be Published Journal: To be PublishedTitle: Solution Structure of (12-53)NCp7 of HIV-1 Authors: Morellet, N. / Demene, H. / Teilleux, V. / Huynh-Dinh, T. / de Rocquigny, H. / Fournie-Zaluski, M.-C. / Roques, B.P. #1:  Journal: J.Mol.Biol. / Year: 1994 Journal: J.Mol.Biol. / Year: 1994Title: Conformational Behaviour of the Active and Inactive Forms of the Nucleocapsid NCp7 of HIV-1 Studied by 1H NMR. Authors: Morellet, N. / de Roquigny, H. / Mely, Y. / Jullian, N. / Demene, H. / Ottmann, M. / Gerard, D. / Darlix, J.L. / Fournie-Zaluski, M.C. / Roques, B.P. #2:  Journal: J.Mol.Biol. / Year: 1998 Journal: J.Mol.Biol. / Year: 1998Title: Structure of the Complex Between the HIV-1 Nucleocapsid Protein and the Single-stranded Pentanucleotide d(ACGCC). Authors: Morellet, N. / Demene, H. / Teilleux, V. / Huynh-Dinh, T. / de Roquigny, H. / Fournie-Zaluski, M.C. / Roques, B.P. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1esk.cif.gz 1esk.cif.gz | 125.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1esk.ent.gz pdb1esk.ent.gz | 101.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1esk.json.gz 1esk.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/es/1esk https://data.pdbj.org/pub/pdb/validation_reports/es/1esk ftp://data.pdbj.org/pub/pdb/validation_reports/es/1esk ftp://data.pdbj.org/pub/pdb/validation_reports/es/1esk | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 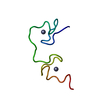
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 4837.642 Da / Num. of mol.: 1 / Fragment: RESIDUES 12-53 / Source method: obtained synthetically Details: The protein was chemically synthesized. The sequence is naturally found in human immunodeficiency virus type 1 (HIV-1). References: UniProt: P04585 |
|---|---|
| #2: Chemical |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||
| NMR details | Text: This structure was determined using standard 2D homonuclear techniques. |
- Sample preparation
Sample preparation
| Details | Contents: 2mM (12-53)NCp7, 90%H2O, 10% D2O, pH 6.0 / Solvent system: 90% H2O/10% D2O |
|---|---|
| Sample conditions | Ionic strength: n.a. / pH: 6 / Pressure: ambient / Temperature: 293 K |
-NMR measurement
| NMR spectrometer | Type: Bruker AMX / Manufacturer: Bruker / Model: AMX / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: simulated annealing / Software ordinal: 1 Details: the structures are based on a total of 444 NOE-derived distance constraints | ||||||||||||
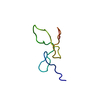
| NMR representative | Selection criteria: lowest energy | ||||||||||||
| NMR ensemble | Conformer selection criteria: structures with acceptable covalent geometry,structures with favorable non-bond energy,structures with the least restraint violations,structures with the lowest energy Conformers calculated total number: 50 / Conformers submitted total number: 9 |
 Movie
Movie Controller
Controller










 PDBj
PDBj









