+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1dzi | ||||||
|---|---|---|---|---|---|---|---|
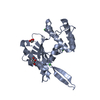
| Title | integrin alpha2 I domain / collagen complex | ||||||
 Components Components |
| ||||||
 Keywords Keywords | INTEGRIN / COLLAGEN | ||||||
| Function / homology |  Function and homology information Function and homology informationcollagen receptor activity / substrate-dependent cell migration / positive regulation of transmission of nerve impulse / response to parathyroid hormone / collagen binding involved in cell-matrix adhesion / integrin alpha2-beta1 complex / hypotonic response / positive regulation of cell projection organization / Regulation of MITF-M-dependent genes involved in extracellular matrix, focal adhesion and epithelial-to-mesenchymal transition / skin morphogenesis ...collagen receptor activity / substrate-dependent cell migration / positive regulation of transmission of nerve impulse / response to parathyroid hormone / collagen binding involved in cell-matrix adhesion / integrin alpha2-beta1 complex / hypotonic response / positive regulation of cell projection organization / Regulation of MITF-M-dependent genes involved in extracellular matrix, focal adhesion and epithelial-to-mesenchymal transition / skin morphogenesis / CHL1 interactions / response to L-ascorbic acid / collagen-activated signaling pathway / positive regulation of smooth muscle contraction / Laminin interactions / basal part of cell / Platelet Adhesion to exposed collagen / hepatocyte differentiation / positive regulation of phagocytosis, engulfment / focal adhesion assembly / mammary gland development / mesodermal cell differentiation / heparan sulfate proteoglycan binding / positive regulation of positive chemotaxis / positive regulation of leukocyte migration / response to amine / integrin complex / cell adhesion mediated by integrin / MET activates PTK2 signaling / positive regulation of smooth muscle cell migration / Syndecan interactions / response to muscle activity / cell-substrate adhesion / positive regulation of collagen biosynthetic process / positive regulation of epithelial cell migration / ECM proteoglycans / Integrin cell surface interactions / detection of mechanical stimulus involved in sensory perception of pain / collagen binding / laminin binding / extracellular matrix organization / positive regulation of cell adhesion / positive regulation of smooth muscle cell proliferation / axon terminus / animal organ morphogenesis / cell-matrix adhesion / integrin-mediated signaling pathway / positive regulation of translation / cellular response to estradiol stimulus / female pregnancy / cellular response to mechanical stimulus / cell-cell adhesion / integrin binding / blood coagulation / amyloid-beta binding / virus receptor activity / signaling receptor activity / response to hypoxia / cell adhesion / response to xenobiotic stimulus / external side of plasma membrane / focal adhesion / protein-containing complex binding / perinuclear region of cytoplasm / cell surface / metal ion binding / plasma membrane Similarity search - Function | ||||||
| Biological species |  HOMO SAPIENS (human) HOMO SAPIENS (human)synthetic construct (others) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.1 Å MOLECULAR REPLACEMENT / Resolution: 2.1 Å | ||||||
 Authors Authors | Emsley, j. / Knight, G. / Farndale, R. / Barnes, M. / Liddington, R. | ||||||
 Citation Citation |  Journal: Cell(Cambridge,Mass.) / Year: 2000 Journal: Cell(Cambridge,Mass.) / Year: 2000Title: Structural Basis of Collagen Recognition by Integrin Alpha2Beta1 Authors: Emsley, J. / Knight, G. / Farndale, R. / Barnes, M. / Liddington, R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1dzi.cif.gz 1dzi.cif.gz | 71.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1dzi.ent.gz pdb1dzi.ent.gz | 52.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1dzi.json.gz 1dzi.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dz/1dzi https://data.pdbj.org/pub/pdb/validation_reports/dz/1dzi ftp://data.pdbj.org/pub/pdb/validation_reports/dz/1dzi ftp://data.pdbj.org/pub/pdb/validation_reports/dz/1dzi | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 20359.053 Da / Num. of mol.: 1 / Fragment: ALPHA2 I DOMAIN Source method: isolated from a genetically manipulated source Source: (gene. exp.)  HOMO SAPIENS (human) / Production host: HOMO SAPIENS (human) / Production host:  | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| #2: Protein/peptide | Mass: 2013.130 Da / Num. of mol.: 3 / Fragment: TRIMERIC GPOGPOGFOGERGPOGPOGPO 21MERIC PEPTIDE / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) #3: Chemical | ChemComp-CO / | #4: Water | ChemComp-HOH / | Has protein modification | Y | Sequence details | CHAIN A: C-TERMINAL ALA IS A SER IN ITA2_HUMAN | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.2 Å3/Da / Density % sol: 44.19 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 6.5 / Details: pH 6.50 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ / pH: 7.5 / Method: vapor diffusion, sitting drop | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SRS SRS  / Beamline: PX7.2 / Wavelength: 1.488 / Beamline: PX7.2 / Wavelength: 1.488 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.488 Å / Relative weight: 1 |
| Reflection | Resolution: 2.1→20 Å / Num. obs: 14483 / % possible obs: 98.2 % / Observed criterion σ(I): 0 / Redundancy: 2.9 % / Rmerge(I) obs: 0.089 / Rsym value: 0.089 / Net I/σ(I): 12 |
| Reflection | *PLUS Lowest resolution: 20 Å |
| Reflection shell | *PLUS Highest resolution: 2.1 Å / Lowest resolution: 2.17 Å / Rmerge(I) obs: 0.344 / Mean I/σ(I) obs: 2.9 |
- Processing
Processing
| Software | Name: CNS / Version: 1 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 2.1→20 Å / Cross valid method: THROUGHOUT / σ(F): 2 / Details: THE RESIDUES B1-3,C1-3,D1-3 ARE IN POOR DENSITY MOLECULAR REPLACEMENT / Resolution: 2.1→20 Å / Cross valid method: THROUGHOUT / σ(F): 2 / Details: THE RESIDUES B1-3,C1-3,D1-3 ARE IN POOR DENSITY
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.1→20 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Version: 1 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Lowest resolution: 20 Å / Rfactor obs: 0.203 / Rfactor Rfree: 0.2729 / Rfactor Rwork: 0.203 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller













 PDBj
PDBj














