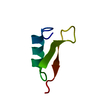
Entry Database : PDB / ID : 1d2jTitle LDL RECEPTOR LIGAND-BINDING MODULE 6 LOW-DENSITY LIPOPROTEIN RECEPTOR Keywords / / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Homo sapiens (human)Method / Authors North, C.L. / Blacklow, S.C. History Deposition Sep 23, 1999 Deposition site / Processing site Revision 1.0 Mar 22, 2000 Provider / Type Revision 1.1 Apr 27, 2008 Group Revision 1.2 Jul 13, 2011 Group Revision 1.3 Nov 3, 2021 Group / Database references / Derived calculationsCategory database_2 / pdbx_nmr_software ... database_2 / pdbx_nmr_software / pdbx_struct_assembly / pdbx_struct_conn_angle / pdbx_struct_oper_list / struct_conn / struct_ref_seq_dif / struct_site Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_nmr_software.name / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_label_seq_id / _struct_ref_seq_dif.details / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id Revision 1.4 Nov 20, 2024 Group / Structure summaryCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / pdbx_entry_details / pdbx_modification_feature
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Homo sapiens (human)
Homo sapiens (human) Authors
Authors Citation
Citation Journal: Biochemistry / Year: 2000
Journal: Biochemistry / Year: 2000 Journal: Biochemistry / Year: 1999
Journal: Biochemistry / Year: 1999 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 1d2j.cif.gz
1d2j.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb1d2j.ent.gz
pdb1d2j.ent.gz PDB format
PDB format 1d2j.json.gz
1d2j.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/d2/1d2j
https://data.pdbj.org/pub/pdb/validation_reports/d2/1d2j ftp://data.pdbj.org/pub/pdb/validation_reports/d2/1d2j
ftp://data.pdbj.org/pub/pdb/validation_reports/d2/1d2j Links
Links Assembly
Assembly
 Components
Components Homo sapiens (human) / Plasmid: PMM-LR6* / Species (production host): Escherichia coli / Production host:
Homo sapiens (human) / Plasmid: PMM-LR6* / Species (production host): Escherichia coli / Production host: 
 Sample preparation
Sample preparation Processing
Processing Movie
Movie Controller
Controller












 PDBj
PDBj













 X-PLOR
X-PLOR