+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1orl | ||||||
|---|---|---|---|---|---|---|---|
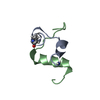
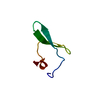
| Title | 1H NMR structure determination of Viscotoxin C1 | ||||||
 Components Components | Viscotoxin C1 | ||||||
 Keywords Keywords | TOXIN / helix-turn-helix / beta-sheet / concentric motif of disulphide bridges | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  Viscum album (European mistletoe) Viscum album (European mistletoe) | ||||||
| Method | SOLUTION NMR / DYANA was employed for structure determination Discover was employed for energy minimisation of DYANA structures | ||||||
 Authors Authors | Molinari, H. / Romagnoli, S. / Fogolari, F. / Catalano, M. / Urech, K. / Giannattasio, M. / Ragona, L. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2003 Journal: Biochemistry / Year: 2003Title: NMR solution structure of viscotoxin C1 from Viscum album species Coloratum ohwi: toward a structure-function analysis of viscotoxins. Authors: Romagnoli, S. / Fogolari, F. / Catalano, M. / Zetta, L. / Schaller, G. / Urech, K. / Giannattasio, M. / Ragona, L. / Molinari, H. #1:  Journal: Biochem.J. / Year: 2000 Journal: Biochem.J. / Year: 2000Title: NMR structural determination of viscotoxin A3 from Viscum album L. Authors: Molinari, H. / Romagnoli, S. / Fogolari, F. / Catalano, M. / Urech, K. / Giannattasio, M. / Ragona, L. | ||||||
| History |
| ||||||
| Remark 999 | sequence The author indicated that the primary sequence for this protein has not yet been deposited ...sequence The author indicated that the primary sequence for this protein has not yet been deposited in any sequence database. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1orl.cif.gz 1orl.cif.gz | 138.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1orl.ent.gz pdb1orl.ent.gz | 113.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1orl.json.gz 1orl.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/or/1orl https://data.pdbj.org/pub/pdb/validation_reports/or/1orl ftp://data.pdbj.org/pub/pdb/validation_reports/or/1orl ftp://data.pdbj.org/pub/pdb/validation_reports/or/1orl | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 4954.694 Da / Num. of mol.: 1 / Source method: isolated from a natural source Details: viscotoxin C1 has been isolated from european mistletoe leaves Source: (natural)  Viscum album (European mistletoe) / References: UniProt: P08943, UniProt: P83554*PLUS Viscum album (European mistletoe) / References: UniProt: P08943, UniProt: P83554*PLUS |
|---|---|
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||||||||||
| NMR details | Text: This structure was determined using standard 1H 2D NMR techniques |
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample conditions |
| ||||||||||||||||||||
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Radiation wavelength | Relative weight: 1 | |||||||||||||||
| NMR spectrometer |
|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: DYANA was employed for structure determination Discover was employed for energy minimisation of DYANA structures Software ordinal: 1 Details: the structures are based on a total of 691 restraints. 631 are NOE-derived distance restraints,(224intraresidues,159 short range, 125 medium range and 123 long range), 18 dihedral angle ...Details: the structures are based on a total of 691 restraints. 631 are NOE-derived distance restraints,(224intraresidues,159 short range, 125 medium range and 123 long range), 18 dihedral angle restraints, 24 for backbone H-bonds and 18 for disulphide bridges. Bond lengths CD-OE2 in glutamic acid residues and CG-OD2 in aspartic acid residues are larger than typical values (as seen in remark 500) because protonated forms were considered due to the low experimental pH. | ||||||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 600 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller








 PDBj
PDBj
