[English] 日本語
 Yorodumi
Yorodumi- PDB-1c2w: 23S RRNA STRUCTURE FITTED TO A CRYO-ELECTRON MICROSCOPIC MAP AT 7... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1c2w | ||||||
|---|---|---|---|---|---|---|---|
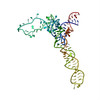
| Title | 23S RRNA STRUCTURE FITTED TO A CRYO-ELECTRON MICROSCOPIC MAP AT 7.5 ANGSTROMS RESOLUTION | ||||||
 Components Components | 23S RIBOSOMAL RNA | ||||||
 Keywords Keywords | RIBOSOME / 23S RRNA / LARGE RIBOSOMAL SUBUNIT / ATOMIC PROTEIN BIOSYNTHESIS / RIBONUCLEIC ACID / EM-RECONSTRUCTION / 3D ARRANGEMENT / FITTING | ||||||
| Function / homology | RNA / RNA (> 10) / RNA (> 100) / RNA (> 1000) Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 7.5 Å | ||||||
 Authors Authors | Brimacombe, R. / Mueller, F. | ||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2000 Journal: J Mol Biol / Year: 2000Title: The 3D arrangement of the 23 S and 5 S rRNA in the Escherichia coli 50 S ribosomal subunit based on a cryo-electron microscopic reconstruction at 7.5 A resolution. Authors: F Mueller / I Sommer / P Baranov / R Matadeen / M Stoldt / J Wöhnert / M Görlach / M van Heel / R Brimacombe /  Abstract: The Escherichia coli 23 S and 5 S rRNA molecules have been fitted helix by helix to a cryo-electron microscopic (EM) reconstruction of the 50 S ribosomal subunit, using an unfiltered version of the ...The Escherichia coli 23 S and 5 S rRNA molecules have been fitted helix by helix to a cryo-electron microscopic (EM) reconstruction of the 50 S ribosomal subunit, using an unfiltered version of the recently published 50 S reconstruction at 7.5 A resolution. At this resolution, the EM density shows a well-defined network of fine structural elements, in which the major and minor grooves of the rRNA helices can be discerned at many locations. The 3D folding of the rRNA molecules within this EM density is constrained by their well-established secondary structures, and further constraints are provided by intra and inter-rRNA crosslinking data, as well as by tertiary interactions and pseudoknots. RNA-protein cross-link and foot-print sites on the 23 S and 5 S rRNA were used to position the rRNA elements concerned in relation to the known arrangement of the ribosomal proteins as determined by immuno-electron microscopy. The published X-ray or NMR structures of seven 50 S ribosomal proteins or RNA-protein complexes were incorporated into the EM density. The 3D locations of cross-link and foot-print sites to the 23 S rRNA from tRNA bound to the ribosomal A, P or E sites were correlated with the positions of the tRNA molecules directly observed in earlier reconstructions of the 70 S ribosome at 13 A or 20 A. Similarly, the positions of cross-link sites within the peptidyl transferase ring of the 23 S rRNA from the aminoacyl residue of tRNA were correlated with the locations of the CCA ends of the A and P site tRNA. Sites on the 23 S rRNA that are cross-linked to the N termini of peptides of different lengths were all found to lie within or close to the internal tunnel connecting the peptidyl transferase region with the presumed peptide exit site on the solvent side of the 50 S subunit. The post-transcriptionally modified bases in the 23 S rRNA form a cluster close to the peptidyl transferase area. The minimum conserved core elements of the secondary structure of the 23 S rRNA form a compact block within the 3D structure and, conversely, the points corresponding to the locations of expansion segments in 28 S rRNA all lie on the outside of the structure. #1:  Journal: J.Mol.Biol. / Year: 1997 Journal: J.Mol.Biol. / Year: 1997Title: A New Model for the Three-Dimensional Folding of Escherichia Coli 16S Ribosomal RNA. I. Fitting the RNA to a 3D Electron Microscopic Map at 20 Angstroms Authors: Mueller, F. / Brimacombe, R. #2:  Journal: J.Mol.Biol. / Year: 1997 Journal: J.Mol.Biol. / Year: 1997Title: A New Model for the Three-Dimensional Folding of Escherichia Coli 16S Ribosomal RNA. II. The RNA-Protein Interaction Data Authors: Mueller, F. / Brimacombe, R. #3:  Journal: J.Mol.Biol. / Year: 1997 Journal: J.Mol.Biol. / Year: 1997Title: A New Model for the Three-Dimensional Folding of Escherichia Coli 16S Ribosomal RNA. III. The Topography of the Functional Centre Authors: Mueller, F. / Stark, H. / Van Heel, M. / Rinke-Appel, J. / Brimacombe, R. #4:  Journal: Structure / Year: 1999 Journal: Structure / Year: 1999Title: The Escherichia coli large ribosomal subunit at 7.5 A resolution Authors: Matadeen, R. / Patwardhan, A. / Gowen, B. / Orlova, E.V. / Pape, T. / Cuff, M. / Mueller, F. / Brimacombe, R. / van Heel, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1c2w.cif.gz 1c2w.cif.gz | 1.5 MB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1c2w.ent.gz pdb1c2w.ent.gz | 1 MB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1c2w.json.gz 1c2w.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/c2/1c2w https://data.pdbj.org/pub/pdb/validation_reports/c2/1c2w ftp://data.pdbj.org/pub/pdb/validation_reports/c2/1c2w ftp://data.pdbj.org/pub/pdb/validation_reports/c2/1c2w | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: RNA chain | Mass: 941718.500 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Details: SECOND OF THE 3 RRNA CHAINS OF THE RIBOSOME / Source: (natural)  |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: 50S Ribosomal Subunit / Type: RIBOSOME |
|---|---|
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Crystal | Description: THE CRYST1 AND SCALE RECORDS ARE MEANINGLESS. |
- Electron microscopy imaging
Electron microscopy imaging
| Microscopy | Model: FEI/PHILIPS CM200FEG / Details: from Structure 1999 citation |
|---|---|
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 38000 X / Nominal defocus max: 2000 nm / Nominal defocus min: 1000 nm |
| Specimen holder | Specimen holder model: GATAN LIQUID NITROGEN |
| Image recording | Film or detector model: GENERIC FILM / Num. of real images: 7 |
| Image scans | Scanner model: IMAGE SCIENCE PATCHWORK DENSITOMETER |
- Processing
Processing
| Software | Name: ERNA-3D / Classification: refinement | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EM software | Name: IMAGIC / Version: 5 / Category: CTF correction | |||||||||||||||||||||
| Particle selection | Num. of particles selected: 16000 / Details: from seven micrographs | |||||||||||||||||||||
| Symmetry | Point symmetry: C1 (asymmetric) | |||||||||||||||||||||
| 3D reconstruction | Resolution: 7.5 Å / Resolution method: FSC 3 SIGMA CUT-OFF / Num. of particles: 16000 / Symmetry type: POINT | |||||||||||||||||||||
| Atomic model building | Space: REAL Details: DETAILS--THIS FILE HAS BEEN GENERATED BY THE USE OF ALL RELEVANT BIOCHEMICAL CONSTRAINTS AND THE CONSTRAINTS GIVEN BY THE ELECTRON DENSITY CONTOUR OF THE RIBOSOME, WHICH WAS DERIVED FROM THE ...Details: DETAILS--THIS FILE HAS BEEN GENERATED BY THE USE OF ALL RELEVANT BIOCHEMICAL CONSTRAINTS AND THE CONSTRAINTS GIVEN BY THE ELECTRON DENSITY CONTOUR OF THE RIBOSOME, WHICH WAS DERIVED FROM THE CRYO-ELECTRON MICROSCOPIC RECONSTRUCTION. THIS FILE IS PART OF A SET OF ALL THREE RIBONUCLEIC ACID CHAINS OF THE RIBOSOME TOGETHER WITH A NUMBER OF RIBOSOMAL PROTEINS WHICH ARE ALSO DEPOSITED WITH THE PDB DATA BANK. THE ATOMIC COORDINATES OF ALL THESE RIBOSOMAL COMPONENTS REFLECT THEIR POSITIONS IN THE 70S RIBOSOME. | |||||||||||||||||||||
| Atomic model building |
| |||||||||||||||||||||
| Refinement | Highest resolution: 7.5 Å / Details: CRYO-EM RECONSTRUCTION | |||||||||||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 7.5 Å
|
 Movie
Movie Controller
Controller













 PDBj
PDBj































