[English] 日本語
 Yorodumi
Yorodumi- PDB-1bhc: BOVINE PANCREATIC TRYPSIN INHIBITOR CRYSTALLIZED FROM THIOCYANATE -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1bhc | ||||||
|---|---|---|---|---|---|---|---|
| Title | BOVINE PANCREATIC TRYPSIN INHIBITOR CRYSTALLIZED FROM THIOCYANATE | ||||||
 Components Components | BOVINE PANCREATIC TRYPSIN INHIBITOR | ||||||
 Keywords Keywords | PROTEASE INHIBITOR / TRYPSIN | ||||||
| Function / homology |  Function and homology information Function and homology informationtrypsinogen activation / negative regulation of serine-type endopeptidase activity / sulfate binding / potassium channel inhibitor activity / negative regulation of platelet aggregation / zymogen binding / molecular function inhibitor activity / negative regulation of thrombin-activated receptor signaling pathway / serine protease inhibitor complex / serine-type endopeptidase inhibitor activity ...trypsinogen activation / negative regulation of serine-type endopeptidase activity / sulfate binding / potassium channel inhibitor activity / negative regulation of platelet aggregation / zymogen binding / molecular function inhibitor activity / negative regulation of thrombin-activated receptor signaling pathway / serine protease inhibitor complex / serine-type endopeptidase inhibitor activity / protease binding / calcium ion binding / extracellular space Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.7 Å MOLECULAR REPLACEMENT / Resolution: 2.7 Å | ||||||
 Authors Authors | Hamiaux, C. / Prange, T. | ||||||
 Citation Citation |  Journal: Acta Crystallogr.,Sect.D / Year: 1999 Journal: Acta Crystallogr.,Sect.D / Year: 1999Title: The decameric structure of bovine pancreatic trypsin inhibitor (BPTI) crystallized from thiocyanate at 2.7 A resolution. Authors: Hamiaux, C. / Prange, T. / Ries-Kautt, M. / Ducruix, A. / Lafont, S. / Astier, J.P. / Veesler, S. #1:  Journal: J.Cryst.Growth / Year: 1997 Journal: J.Cryst.Growth / Year: 1997Title: Comparison of Solubilities and Molecular Interactions of Bpti Molecules Giving Different Polymorphs Authors: Lafont, S. / Veesler, S. / Astier, J.P. / Boistelle, R. #2:  Journal: Acta Crystallogr.,Sect.D / Year: 1995 Journal: Acta Crystallogr.,Sect.D / Year: 1995Title: Structure of Hexagonal Turkey Egg-White Lysozyme at 1.65 A Resolution Authors: Howell, P.L. #3:  Journal: Acta Crystallogr.,Sect.B / Year: 1992 Journal: Acta Crystallogr.,Sect.B / Year: 1992Title: Structure Determination of a Dimeric Form of Erabutoxin-B, Crystallized from a Thiocyanate Solution Authors: Saludjian, P. / Prange, T. / Navaza, J. / Menez, R. / Guilloteau, J.P. / Ries-Kautt, M. / Ducruix, A. #4:  Journal: J.Mol.Biol. / Year: 1987 Journal: J.Mol.Biol. / Year: 1987Title: Comparison of Two Highly Refined Structures of Bovine Pancreatic Trypsin Inhibitor Authors: Wlodawer, A. / Deisenhofer, J. / Huber, R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1bhc.cif.gz 1bhc.cif.gz | 122.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1bhc.ent.gz pdb1bhc.ent.gz | 98.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1bhc.json.gz 1bhc.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1bhc_validation.pdf.gz 1bhc_validation.pdf.gz | 424.3 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1bhc_full_validation.pdf.gz 1bhc_full_validation.pdf.gz | 436.2 KB | Display | |
| Data in XML |  1bhc_validation.xml.gz 1bhc_validation.xml.gz | 12.2 KB | Display | |
| Data in CIF |  1bhc_validation.cif.gz 1bhc_validation.cif.gz | 20.2 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/bh/1bhc https://data.pdbj.org/pub/pdb/validation_reports/bh/1bhc ftp://data.pdbj.org/pub/pdb/validation_reports/bh/1bhc ftp://data.pdbj.org/pub/pdb/validation_reports/bh/1bhc | HTTPS FTP |
-Related structure data
| Related structure data | 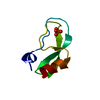 6ptiS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||
| Unit cell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||
| Noncrystallographic symmetry (NCS) | NCS domain:
NCS oper:
|
- Components
Components
| #1: Protein | Mass: 6527.568 Da / Num. of mol.: 10 / Source method: isolated from a natural source / Source: (natural)  #2: Chemical | ChemComp-SCN / #3: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.61 Å3/Da / Density % sol: 52 % |
|---|---|
| Crystal grow | pH: 4.5 Details: BPTI WAS CRYSTALLIZED FROM 250MM THIOCYANATE IN ACETATE BUFFER (50MM, PH=4.5) |
| Crystal grow | *PLUS Method: other / Details: Lafont, S., (1997) J. Crystal Growth, 173, 132. |
-Data collection
| Diffraction | Mean temperature: 292 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: LURE SYNCHROTRON / Site: LURE  / Beamline: DW32 / Wavelength: 0.97 / Beamline: DW32 / Wavelength: 0.97 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Mar 1, 1997 / Details: MIRRORS |
| Radiation | Monochromator: SI(111) / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.97 Å / Relative weight: 1 |
| Reflection | Resolution: 2.63→29.59 Å / Num. obs: 18308 / % possible obs: 85.5 % / Observed criterion σ(I): 2 / Redundancy: 3.8 % / Biso Wilson estimate: 50 Å2 / Rmerge(I) obs: 0.055 / Rsym value: 0.055 / Net I/σ(I): 11.1 |
| Reflection shell | Resolution: 2.63→2.71 Å / Redundancy: 3.4 % / Rmerge(I) obs: 0.186 / Mean I/σ(I) obs: 4.1 / Rsym value: 0.186 / % possible all: 13.4 |
| Reflection | *PLUS Num. measured all: 69959 |
| Reflection shell | *PLUS % possible obs: 13.4 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 6PTI Resolution: 2.7→10 Å / Data cutoff high absF: 10000000 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 2 Details: RESIDUES ARG1, ASP3, LYS15, LYS26 AND ARG39 APPEAR TO HAVE NCS BREAKDOWN IN RELATED MOLECULES. THEY WERE REMOVED FROM THE NCS RESTRAINT SCHEME. MET 52 WAS MODELLED WITH TWO CONFORMATIONS IN ...Details: RESIDUES ARG1, ASP3, LYS15, LYS26 AND ARG39 APPEAR TO HAVE NCS BREAKDOWN IN RELATED MOLECULES. THEY WERE REMOVED FROM THE NCS RESTRAINT SCHEME. MET 52 WAS MODELLED WITH TWO CONFORMATIONS IN ALL MOLECULES. 10 THIOCYANATE IONS AND 118 WATER MOLECULES ARE GIVEN FOLLOWING THE COORDINATES OF THE TEN MOLECULES. AS IN 6PTI, NO DENSITY WAS OBSERVED FOR THE TWO LAST RESIDUES (GLY 57 & ALA 58).
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 25.65 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.7→10 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints NCS | Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.7→2.82 Å / Total num. of bins used: 8
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.01 / Classification: refinement X-PLOR / Version: 3.01 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor Rfree: 0.4 |
 Movie
Movie Controller
Controller









 PDBj
PDBj







