+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1b9a | ||||||
|---|---|---|---|---|---|---|---|
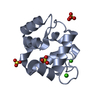
| Title | PARVALBUMIN (MUTATION;D51A, F102W) | ||||||
 Components Components | PROTEIN (PARVALBUMIN) | ||||||
 Keywords Keywords | CALCIUM BINDING PROTEIN / EF-HAND PROTEINS / PARVALBUMIN / CALCIUM-BINDING | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  Cyprinus carpio (common carp) Cyprinus carpio (common carp) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2 Å MOLECULAR REPLACEMENT / Resolution: 2 Å | ||||||
 Authors Authors | Cates, M.S. / Berry, M.B. / Ho, E.L. / Li, Q. / Potter, J.D. / Phillips Jr., G.N. | ||||||
 Citation Citation |  Journal: Structure Fold.Des. / Year: 1999 Journal: Structure Fold.Des. / Year: 1999Title: Metal-ion affinity and specificity in EF-hand proteins: coordination geometry and domain plasticity in parvalbumin. Authors: Cates, M.S. / Berry, M.B. / Ho, E.L. / Li, Q. / Potter, J.D. / Phillips Jr., G.N. #1:  Journal: J.Biol.Chem. / Year: 1989 Journal: J.Biol.Chem. / Year: 1989Title: Restrained Least Squares Refinement of Native (Calcium) and Cadmium- Substituted Carp Parvalbumin Using X-Ray Crystallographic Data at 1.6 Angstrom Resolution Authors: Swain, A.L. / Kretsinger, R.H. / Amma, E.L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1b9a.cif.gz 1b9a.cif.gz | 37.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1b9a.ent.gz pdb1b9a.ent.gz | 25.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1b9a.json.gz 1b9a.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/b9/1b9a https://data.pdbj.org/pub/pdb/validation_reports/b9/1b9a ftp://data.pdbj.org/pub/pdb/validation_reports/b9/1b9a ftp://data.pdbj.org/pub/pdb/validation_reports/b9/1b9a | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1b8cC  1b8lC  1b8rC  5cpvS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||
| Unit cell |
| ||||||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 11445.816 Da / Num. of mol.: 1 / Mutation: D51A, F102W Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Cyprinus carpio (common carp) / Production host: Cyprinus carpio (common carp) / Production host:  | ||
|---|---|---|---|
| #2: Chemical | | #3: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.1 Å3/Da / Density % sol: 41.44 % | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 7 / Details: pH 7.0 | ||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ / pH: 7 / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 110 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X4A / Wavelength: 1.743318 / Beamline: X4A / Wavelength: 1.743318 |
| Detector | Type: FUJI / Detector: IMAGE PLATE / Date: Jun 1, 1997 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.743318 Å / Relative weight: 1 |
| Reflection | Resolution: 2→40 Å / Num. obs: 5432 / % possible obs: 73 % / Biso Wilson estimate: 14.9 Å2 / Rmerge(I) obs: 0.069 |
| Reflection | *PLUS % possible obs: 73 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 5CPV, AUTH A.L.SWAIN,R.H.KRETSINGER, E.L.AMMA Resolution: 2→40 Å / Rfactor Rfree error: 0.012 / Data cutoff high rms absF: 723489.75 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 37.41 Å2 / ksol: 0.3181 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 23.1 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→40 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2→2.13 Å / Rfactor Rfree error: 0.047 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: CNS / Version: 0.4 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS σ(F): 0 / Rfactor obs: 0.2096 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS Biso mean: 23.1 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor Rfree: 0.403 / % reflection Rfree: 12.2 % / Rfactor Rwork: 0.275 |
 Movie
Movie Controller
Controller












 PDBj
PDBj


