[English] 日本語
 Yorodumi
Yorodumi- PDB-1ayb: CRYSTAL STRUCTURES OF PEPTIDE COMPLEXES OF THE AMINO-TERMINAL SH2... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ayb | ||||||
|---|---|---|---|---|---|---|---|
| Title | CRYSTAL STRUCTURES OF PEPTIDE COMPLEXES OF THE AMINO-TERMINAL SH2 DOMAIN OF THE SYP TYROSINE PHOSPHATASE | ||||||
 Components Components |
| ||||||
 Keywords Keywords | HYDROLASE(SH2 DOMAIN) | ||||||
| Function / homology |  Function and homology information Function and homology informationIRS-mediated signalling / Signaling by ALK / IRS-related events triggered by IGF1R / PI-3K cascade:FGFR3 / PI-3K cascade:FGFR4 / Co-inhibition by BTLA / PI-3K cascade:FGFR1 / PI-3K cascade:FGFR2 / SOS-mediated signalling / FRS-mediated FGFR3 signaling ...IRS-mediated signalling / Signaling by ALK / IRS-related events triggered by IGF1R / PI-3K cascade:FGFR3 / PI-3K cascade:FGFR4 / Co-inhibition by BTLA / PI-3K cascade:FGFR1 / PI-3K cascade:FGFR2 / SOS-mediated signalling / FRS-mediated FGFR3 signaling / FRS-mediated FGFR4 signaling / MET activates PTPN11 / Regulation of IFNA/IFNB signaling / Signaling by LTK / FRS-mediated FGFR1 signaling / FRS-mediated FGFR2 signaling / Tie2 Signaling / PI3K/AKT activation / PECAM1 interactions / Interleukin-20 family signaling / Signaling by CSF3 (G-CSF) / GAB1 signalosome / PI3K Cascade / MAPK3 (ERK1) activation / MAPK1 (ERK2) activation / Co-inhibition by CTLA4 / Regulation of RUNX1 Expression and Activity / negative regulation of somatostatin secretion / Interleukin-6 signaling / Downstream signal transduction / positive regulation of glucagon secretion / GPVI-mediated activation cascade / Interleukin-7 signaling / IRS activation / Co-inhibition by PD-1 / epithelial cell migration / Signal attenuation / Signaling by SCF-KIT / Negative regulation of FGFR3 signaling / Negative regulation of FGFR4 signaling / Negative regulation of FGFR1 signaling / Negative regulation of FGFR2 signaling / Spry regulation of FGF signaling / Activation of IRF3, IRF7 mediated by TBK1, IKKε (IKBKE) / positive regulation of signal transduction / negative regulation of hormone secretion / PIP3 activates AKT signaling / negative regulation of cortisol secretion / intestinal epithelial cell migration / microvillus organization / negative regulation of growth hormone secretion / RAF/MAP kinase cascade / Platelet sensitization by LDL / genitalia development / atrioventricular canal development / PI5P, PP2A and IER3 Regulate PI3K/AKT Signaling / positive regulation of fatty acid beta-oxidation / positive regulation of glucose metabolic process / Interleukin-3, Interleukin-5 and GM-CSF signaling / RET signaling / phosphatidylinositol 3-kinase activator activity / negative regulation of neutrophil activation / cerebellar cortex formation / insulin receptor complex / positive regulation of hormone secretion / transmembrane receptor protein tyrosine kinase adaptor activity / mammary gland development / regulation of protein export from nucleus / positive regulation of lipopolysaccharide-mediated signaling pathway / positive regulation of ossification / negative regulation of cell adhesion mediated by integrin / cellular response to fatty acid / regulation of cell adhesion mediated by integrin / hormone metabolic process / negative regulation of chondrocyte differentiation / response to caffeine / face morphogenesis / positive regulation of mesenchymal cell proliferation / organ growth / platelet formation / ERBB signaling pathway / triglyceride metabolic process / megakaryocyte development / negative regulation of T cell activation / peptide hormone receptor binding / negative regulation of type I interferon production / neurotrophin TRK receptor signaling pathway / platelet-derived growth factor receptor signaling pathway / inner ear development / Bergmann glial cell differentiation / non-membrane spanning protein tyrosine phosphatase activity / regulation of MAPK cascade / positive regulation of intracellular signal transduction / positive regulation of epithelial cell migration / insulin receptor substrate binding / fibroblast growth factor receptor signaling pathway / positive regulation of phosphorylation / positive regulation of focal adhesion assembly / plasma membrane raft / positive regulation of glycogen biosynthetic process Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 3 Å X-RAY DIFFRACTION / Resolution: 3 Å | ||||||
 Authors Authors | Lee, C.-H. / Kuriyan, J. | ||||||
 Citation Citation |  Journal: Structure / Year: 1994 Journal: Structure / Year: 1994Title: Crystal structures of peptide complexes of the amino-terminal SH2 domain of the Syp tyrosine phosphatase. Authors: Lee, C.H. / Kominos, D. / Jacques, S. / Margolis, B. / Schlessinger, J. / Shoelson, S.E. / Kuriyan, J. #1:  Journal: Cell(Cambridge,Mass.) / Year: 1993 Journal: Cell(Cambridge,Mass.) / Year: 1993Title: Binding of a High Affinity Phosphotyrosyl Peptide to the Src Sh2 Domain: Crystal Structures of the Complexed and Peptide-Free Forms Authors: Waksman, G. / Shoelson, S.E. / Pant, N. / Cowburn, D. / Kuriyan, J. #2:  Journal: Curr.Opin.Struct.Biol. / Year: 1993 Journal: Curr.Opin.Struct.Biol. / Year: 1993Title: Structures of Sh2 and SH3 Domains Authors: Kuriyan, J. / Cowburn, D. #3:  Journal: Nature / Year: 1992 Journal: Nature / Year: 1992Title: Crystal Structure of the Phosphotyrosine Recognition Domain Sh2 of V-Src Complexed with Tyrosine-Phosphorylated Peptides Authors: Waksman, G. / Kominos, D. / Robertson, S.R. / Pant, N. / Baltimore, D. / Birge, R.B. / Cowburn, D. / Hanafusa, H. / Mayer, B.J. / Overduin, M. / Resh, M.D. / Rios, C.B. / Silverman, L. / Kuriyan, J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ayb.cif.gz 1ayb.cif.gz | 34.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ayb.ent.gz pdb1ayb.ent.gz | 21.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ayb.json.gz 1ayb.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ay/1ayb https://data.pdbj.org/pub/pdb/validation_reports/ay/1ayb ftp://data.pdbj.org/pub/pdb/validation_reports/ay/1ayb ftp://data.pdbj.org/pub/pdb/validation_reports/ay/1ayb | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 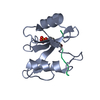
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 11528.979 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  |
|---|---|
| #2: Protein/peptide | Mass: 1378.335 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.98 Å3/Da / Density % sol: 58.72 % | ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | *PLUS Temperature: 4 ℃ / pH: 6 / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Radiation | Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| Reflection | *PLUS Highest resolution: 3 Å / Lowest resolution: 30 Å / Num. all: 5965 / Num. obs: 2549 / % possible obs: 73 % / Rmerge(I) obs: 0.089 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 3→6 Å / σ(F): 2 /
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 3→6 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.164 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS Type: x_angle_d / Dev ideal: 3 |
 Movie
Movie Controller
Controller












 PDBj
PDBj









