+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1a7v | ||||||
|---|---|---|---|---|---|---|---|
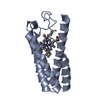
| Title | CYTOCHROME C' FROM RHODOPSEUDOMONAS PALUSTRIS | ||||||
 Components Components | CYTOCHROME C' | ||||||
 Keywords Keywords | ELECTRON TRANSPORT | ||||||
| Function / homology |  Function and homology information Function and homology informationelectron transport chain / periplasmic space / electron transfer activity / iron ion binding / heme binding Similarity search - Function | ||||||
| Biological species |  Rhodopseudomonas palustris (phototrophic) Rhodopseudomonas palustris (phototrophic) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.3 Å MOLECULAR REPLACEMENT / Resolution: 2.3 Å | ||||||
 Authors Authors | Shibata, N. / Iba, S. / Misaki, S. / Meyer, T.E. / Bartsch, R.G. / Cusanovich, M.A. / Higuchi, Y. / Yasuoka, N. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1998 Journal: J.Mol.Biol. / Year: 1998Title: Basis for monomer stabilization in Rhodopseudomonas palustris cytochrome c' derived from the crystal structure. Authors: Shibata, N. / Iba, S. / Misaki, S. / Meyer, T.E. / Bartsch, R.G. / Cusanovich, M.A. / Morimoto, Y. / Higuchi, Y. / Yasuoka, N. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1a7v.cif.gz 1a7v.cif.gz | 64.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1a7v.ent.gz pdb1a7v.ent.gz | 47.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1a7v.json.gz 1a7v.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/a7/1a7v https://data.pdbj.org/pub/pdb/validation_reports/a7/1a7v ftp://data.pdbj.org/pub/pdb/validation_reports/a7/1a7v ftp://data.pdbj.org/pub/pdb/validation_reports/a7/1a7v | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2ccyS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (-0.997291, 0.046599, 0.056917), Vector: |
- Components
Components
| #1: Protein | Mass: 13152.139 Da / Num. of mol.: 2 / Source method: isolated from a natural source / Source: (natural)  Rhodopseudomonas palustris (phototrophic) / References: UniProt: P00149 Rhodopseudomonas palustris (phototrophic) / References: UniProt: P00149#2: Chemical | #3: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.23 Å3/Da / Density % sol: 45 % | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 6 / Details: pH 6.0 | ||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion, sitting dropDetails: drop solution was mixed with an equal volume of reservoir solution | ||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 293 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Photon Factory Photon Factory  / Beamline: BL-6A / Wavelength: 1 / Beamline: BL-6A / Wavelength: 1 |
| Detector | Detector: IMAGE PLATE / Date: Jul 1, 1996 |
| Radiation | Monochromator: SI(111) / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 2.3→30 Å / Num. obs: 10513 / % possible obs: 89.1 % / Observed criterion σ(I): 1 / Redundancy: 3.7 % / Biso Wilson estimate: 17.1 Å2 / Rmerge(I) obs: 0.072 / Net I/σ(I): 15.9 |
| Reflection shell | Resolution: 2.3→2.38 Å / Redundancy: 2.1 % / Rmerge(I) obs: 0.16 / Mean I/σ(I) obs: 2.1 / % possible all: 63 |
| Reflection | *PLUS Num. measured all: 39136 |
| Reflection shell | *PLUS % possible obs: 62.7 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 2CCY Resolution: 2.3→7 Å / Data cutoff high absF: 100000 / Data cutoff low absF: 0.1 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.25 Å / Luzzati d res low obs: 7 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.3→7 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints NCS | NCS model details: RESTRAINED / Rms dev Biso : 2 Å2 / Rms dev position: 0.1 Å / Weight Biso : 4.8 / Weight position: 100 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.3→2.4 Å / Total num. of bins used: 8
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.1 / Classification: refinement X-PLOR / Version: 3.1 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor obs: 0.243 |
 Movie
Movie Controller
Controller











 PDBj
PDBj








