+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: PDB / ID: 136d | ||||||
|---|---|---|---|---|---|---|---|
| タイトル | SOLUTION STRUCTURE OF A PURINE(DOT)PURINE(DOT)PYRIMIDINE DNA TRIPLEX CONTAINING G(DOT)GC AND T(DOT)AT TRIPLES | ||||||
 要素 要素 | DNA TRIPLEX | ||||||
 キーワード キーワード | DNA / TRIPLEX | ||||||
| 機能・相同性 | DNA / DNA (> 10) 機能・相同性情報 機能・相同性情報 | ||||||
| 手法 | 溶液NMR / RESTRAINED MOLECULAR DYNAMICS, DISTANCE GEOMETRY | ||||||
 データ登録者 データ登録者 | Radhakrishnan, I. / Patel, D.J. | ||||||
 引用 引用 |  ジャーナル: Structure / 年: 1993 ジャーナル: Structure / 年: 1993タイトル: Solution structure of a purine.purine.pyrimidine DNA triplex containing G.GC and T.AT triples. 著者: Radhakrishnan, I. / Patel, D.J. #1:  ジャーナル: J.Am.Chem.Soc. / 年: 1993 ジャーナル: J.Am.Chem.Soc. / 年: 1993タイトル: Solution Structure of an Intramolecular Purine(Dot)Purine(Dot)Pyrimidine DNA Triplex 著者: Radhakrishnan, I. / Patel, D.J. | ||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 構造ビューア | 分子:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- ダウンロードとリンク
ダウンロードとリンク
- ダウンロード
ダウンロード
| PDBx/mmCIF形式 |  136d.cif.gz 136d.cif.gz | 261.2 KB | 表示 |  PDBx/mmCIF形式 PDBx/mmCIF形式 |
|---|---|---|---|---|
| PDB形式 |  pdb136d.ent.gz pdb136d.ent.gz | 181.9 KB | 表示 |  PDB形式 PDB形式 |
| PDBx/mmJSON形式 |  136d.json.gz 136d.json.gz | ツリー表示 |  PDBx/mmJSON形式 PDBx/mmJSON形式 | |
| その他 |  その他のダウンロード その他のダウンロード |
-検証レポート
| 文書・要旨 |  136d_validation.pdf.gz 136d_validation.pdf.gz | 309.1 KB | 表示 |  wwPDB検証レポート wwPDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  136d_full_validation.pdf.gz 136d_full_validation.pdf.gz | 514.1 KB | 表示 | |
| XML形式データ |  136d_validation.xml.gz 136d_validation.xml.gz | 46.4 KB | 表示 | |
| CIF形式データ |  136d_validation.cif.gz 136d_validation.cif.gz | 60.6 KB | 表示 | |
| アーカイブディレクトリ |  https://data.pdbj.org/pub/pdb/validation_reports/36/136d https://data.pdbj.org/pub/pdb/validation_reports/36/136d ftp://data.pdbj.org/pub/pdb/validation_reports/36/136d ftp://data.pdbj.org/pub/pdb/validation_reports/36/136d | HTTPS FTP |
-関連構造データ
- リンク
リンク
- 集合体
集合体
| 登録構造単位 | 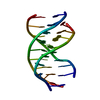
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR アンサンブル |
|
- 要素
要素
| #1: DNA鎖 | 分子量: 9552.122 Da / 分子数: 1 / 由来タイプ: 合成 / 詳細: CHEMICALLY SYNTHESIZED |
|---|
-実験情報
-実験
| 実験 | 手法: 溶液NMR |
|---|---|
| NMR実験の詳細 | Text: STRUCTURE ANALYSES WERE CONDUCTED ON SIXTEEN STRUCTURES INCLUDING FOURTEEN STRUCTURES TAKEN FROM THE RESTRAINED MOLECULAR DYNAMICS TRAJECTORIES OF THE TWO REFINEMENTS, AND THE TWO AVERAGED ...Text: STRUCTURE ANALYSES WERE CONDUCTED ON SIXTEEN STRUCTURES INCLUDING FOURTEEN STRUCTURES TAKEN FROM THE RESTRAINED MOLECULAR DYNAMICS TRAJECTORIES OF THE TWO REFINEMENTS, AND THE TWO AVERAGED MINIMIZED STRUCTURES. THE COORDINATES FOR THE TWO AVERAGED MINIMIZED STRUCTURES CAN BE FOUND IN PROTEIN DATA BANK ENTRIES 134D AND 135D, RESPECTIVELY. THE COORDINATES FOR THE 14 STRUCTURES FROM THE RESTRAINED MOLECULAR DYNAMICS TRAJECTORIES CAN BE FOUND IN ENTRY 136D. |
- 解析
解析
| ソフトウェア |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| NMR software | 名称:  X-PLOR / 開発者: BRUNGER / 分類: 精密化 X-PLOR / 開発者: BRUNGER / 分類: 精密化 | |||||||||
| 精密化 | 手法: RESTRAINED MOLECULAR DYNAMICS, DISTANCE GEOMETRY / ソフトェア番号: 1 詳細: RESTRAINED MOLECULAR DYNAMICS CALCULATIONS WERE DONE ON IDEALIZED A'- AND B-FORM STARTING STRUCTURES. ONLY THE TRIPLEX REGION AND THE FIRST AND LAST RESIDUES OF EACH OF THE TWO LOOPS IN THE ...詳細: RESTRAINED MOLECULAR DYNAMICS CALCULATIONS WERE DONE ON IDEALIZED A'- AND B-FORM STARTING STRUCTURES. ONLY THE TRIPLEX REGION AND THE FIRST AND LAST RESIDUES OF EACH OF THE TWO LOOPS IN THE SEQUENCE WERE CONSIDERED. THE REFINEMENT WAS CONDUCTED IN TWO STAGES. IN THE FIRST STAGE, SIX STRUCTURES WERE CALCULATED (THREE FROM EACH STARTING STRUCTURE) USING DISTANCE RESTRAINTS. IN THE SECOND STAGE, TWO OF THE SIX STRUCTURES WERE REFINED DIRECTLY AGAINST PRIMARY NOE DATA. THE R(1/6) VALUE WAS USED TO MONITOR THE REFINEMENT DURING THIS STAGE. THE FINAL R(1/6) VALUES FOR THE TWO AVERAGED MINIMIZED STRUCTURES WERE 0.04 AND 0.045. THE SUMMATION RUNS THROUGH ALL OBSERVED, QUANTIFIABLE CROSSPEAK INTENSITIES IN NOESY SPECTRA RECORDED AT MIXING TIMES OF 40, 90 AND 150 MS. RMSD BOND DISTANCES 0.016 ANGSTROMS RMSD BOND ANGLES 3.96 DEGREES RMS DEVIATIONS FROM IDEALIZED GEOMETRY FOR THE TWO AVERAGED MINIMIZED STRUCTURES ARE AS FOLLOWS: | |||||||||
| NMRアンサンブル | 登録したコンフォーマーの数: 14 | |||||||||
| ソフトウェア | *PLUS 名称:  X-PLOR / 分類: refinement X-PLOR / 分類: refinement | |||||||||
| 拘束条件 | *PLUS
|
 ムービー
ムービー コントローラー
コントローラー







 PDBj
PDBj






































