[English] 日本語
 Yorodumi
Yorodumi- PDB-136d: SOLUTION STRUCTURE OF A PURINE(DOT)PURINE(DOT)PYRIMIDINE DNA TRIP... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 136d | ||||||
|---|---|---|---|---|---|---|---|
| Title | SOLUTION STRUCTURE OF A PURINE(DOT)PURINE(DOT)PYRIMIDINE DNA TRIPLEX CONTAINING G(DOT)GC AND T(DOT)AT TRIPLES | ||||||
 Components Components | DNA TRIPLEX | ||||||
 Keywords Keywords | DNA / TRIPLEX | ||||||
| Function / homology | DNA / DNA (> 10) Function and homology information Function and homology information | ||||||
| Method | SOLUTION NMR / RESTRAINED MOLECULAR DYNAMICS, DISTANCE GEOMETRY | ||||||
 Authors Authors | Radhakrishnan, I. / Patel, D.J. | ||||||
 Citation Citation |  Journal: Structure / Year: 1993 Journal: Structure / Year: 1993Title: Solution structure of a purine.purine.pyrimidine DNA triplex containing G.GC and T.AT triples. Authors: Radhakrishnan, I. / Patel, D.J. #1:  Journal: J.Am.Chem.Soc. / Year: 1993 Journal: J.Am.Chem.Soc. / Year: 1993Title: Solution Structure of an Intramolecular Purine(Dot)Purine(Dot)Pyrimidine DNA Triplex Authors: Radhakrishnan, I. / Patel, D.J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  136d.cif.gz 136d.cif.gz | 261.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb136d.ent.gz pdb136d.ent.gz | 181.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  136d.json.gz 136d.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/36/136d https://data.pdbj.org/pub/pdb/validation_reports/36/136d ftp://data.pdbj.org/pub/pdb/validation_reports/36/136d ftp://data.pdbj.org/pub/pdb/validation_reports/36/136d | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 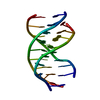
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: DNA chain | Mass: 9552.122 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: CHEMICALLY SYNTHESIZED |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR |
|---|---|
| NMR details | Text: STRUCTURE ANALYSES WERE CONDUCTED ON SIXTEEN STRUCTURES INCLUDING FOURTEEN STRUCTURES TAKEN FROM THE RESTRAINED MOLECULAR DYNAMICS TRAJECTORIES OF THE TWO REFINEMENTS, AND THE TWO AVERAGED ...Text: STRUCTURE ANALYSES WERE CONDUCTED ON SIXTEEN STRUCTURES INCLUDING FOURTEEN STRUCTURES TAKEN FROM THE RESTRAINED MOLECULAR DYNAMICS TRAJECTORIES OF THE TWO REFINEMENTS, AND THE TWO AVERAGED MINIMIZED STRUCTURES. THE COORDINATES FOR THE TWO AVERAGED MINIMIZED STRUCTURES CAN BE FOUND IN PROTEIN DATA BANK ENTRIES 134D AND 135D, RESPECTIVELY. THE COORDINATES FOR THE 14 STRUCTURES FROM THE RESTRAINED MOLECULAR DYNAMICS TRAJECTORIES CAN BE FOUND IN ENTRY 136D. |
- Processing
Processing
| Software |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| NMR software | Name:  X-PLOR / Developer: BRUNGER / Classification: refinement X-PLOR / Developer: BRUNGER / Classification: refinement | |||||||||
| Refinement | Method: RESTRAINED MOLECULAR DYNAMICS, DISTANCE GEOMETRY / Software ordinal: 1 Details: RESTRAINED MOLECULAR DYNAMICS CALCULATIONS WERE DONE ON IDEALIZED A'- AND B-FORM STARTING STRUCTURES. ONLY THE TRIPLEX REGION AND THE FIRST AND LAST RESIDUES OF EACH OF THE TWO LOOPS IN THE ...Details: RESTRAINED MOLECULAR DYNAMICS CALCULATIONS WERE DONE ON IDEALIZED A'- AND B-FORM STARTING STRUCTURES. ONLY THE TRIPLEX REGION AND THE FIRST AND LAST RESIDUES OF EACH OF THE TWO LOOPS IN THE SEQUENCE WERE CONSIDERED. THE REFINEMENT WAS CONDUCTED IN TWO STAGES. IN THE FIRST STAGE, SIX STRUCTURES WERE CALCULATED (THREE FROM EACH STARTING STRUCTURE) USING DISTANCE RESTRAINTS. IN THE SECOND STAGE, TWO OF THE SIX STRUCTURES WERE REFINED DIRECTLY AGAINST PRIMARY NOE DATA. THE R(1/6) VALUE WAS USED TO MONITOR THE REFINEMENT DURING THIS STAGE. THE FINAL R(1/6) VALUES FOR THE TWO AVERAGED MINIMIZED STRUCTURES WERE 0.04 AND 0.045. THE SUMMATION RUNS THROUGH ALL OBSERVED, QUANTIFIABLE CROSSPEAK INTENSITIES IN NOESY SPECTRA RECORDED AT MIXING TIMES OF 40, 90 AND 150 MS. RMSD BOND DISTANCES 0.016 ANGSTROMS RMSD BOND ANGLES 3.96 DEGREES RMS DEVIATIONS FROM IDEALIZED GEOMETRY FOR THE TWO AVERAGED MINIMIZED STRUCTURES ARE AS FOLLOWS: | |||||||||
| NMR ensemble | Conformers submitted total number: 14 | |||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | |||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller






 PDBj
PDBj






































