[English] 日本語
 Yorodumi
Yorodumi- PDB-105d: SOLUTION STRUCTURES OF THE I-MOTIF TETRAMERS OF D(TCC), D(5MCCT) ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 105d | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | SOLUTION STRUCTURES OF THE I-MOTIF TETRAMERS OF D(TCC), D(5MCCT) AND D(T5MCC). NOVEL NOE CONNECTIONS BETWEEN AMINO PROTONS AND SUGAR PROTONS | ||||||||||||||||||
 Components Components | DNA (5'-D(* Keywords KeywordsDNA | Function / homology | DNA |  Function and homology information Function and homology informationMethod | SOLUTION NMR |  Authors AuthorsLeroy, J.-L. / Gueron, M. |  Citation Citation Journal: Structure / Year: 1995 Journal: Structure / Year: 1995Title: Solution structures of the i-motif tetramers of d(TCC), d(5methylCCT) and d(T5methylCC): novel NOE connections between amino protons and sugar protons. Authors: Leroy, J.L. / Gueron, M. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  105d.cif.gz 105d.cif.gz | 55.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb105d.ent.gz pdb105d.ent.gz | 38.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  105d.json.gz 105d.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/05/105d https://data.pdbj.org/pub/pdb/validation_reports/05/105d ftp://data.pdbj.org/pub/pdb/validation_reports/05/105d ftp://data.pdbj.org/pub/pdb/validation_reports/05/105d | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 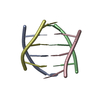
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: DNA chain | Mass: 837.599 Da / Num. of mol.: 4 / Source method: obtained synthetically |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR |
|---|
- Sample preparation
Sample preparation
| Crystal grow | *PLUS Method: other / Details: NMR |
|---|
- Processing
Processing
| Software |
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Refinement | Software ordinal: 1 Details: THE STRUCTURE IS FORMED OF FOUR EQUIVALENT TCC STRANDS DESIGNATED A, B, C, D. STRANDS A AND C, CONNECTED BY TWO HEMI-PROTONATED BASE PAIRS (C2.C2+ AND C3.C3+), FORM A PARALLEL-STRANDED ...Details: THE STRUCTURE IS FORMED OF FOUR EQUIVALENT TCC STRANDS DESIGNATED A, B, C, D. STRANDS A AND C, CONNECTED BY TWO HEMI-PROTONATED BASE PAIRS (C2.C2+ AND C3.C3+), FORM A PARALLEL-STRANDED DUPLEX. THE RELATION BETWEEN STRANDS B AND D IS THE SAME AS THAT BETWEEN STRANDS A AND C. THE STRANDS OF EACH DUPLEX ARE RELATED BY A LONGITUDINAL TWO-FOLD SYMMETRY AXIS. THE DUPLEXES HAVE THE SAME LONGITUDINAL AXIS, AND THEY ARE ANTI-PARALLEL. THEIR HEMI-PROTONATED C.C+ BASE PAIRS ARE INTERCALATED FACE-TO-FACE. THE DUPLEXES CAN BE TRANSFORMED INTO ONE ANOTHER BY A 180 DEGREE ROTATION AROUND A TWO-FOLD AXIS PERPENDICULAR TO THE LONGITUDINAL AXIS, AND CONSEQUENTLY AROUND A THIRD AXIS PERPENDICULAR TO THE FIRST TWO. THE H3 IMINO PROTONS OF THE HEMI-PROTONATED C.C+ PAIRS WERE NOT INCORPORATED IN THE COMPUTATIONS. | ||||||||
| NMR ensemble | Conformers submitted total number: 8 |
 Movie
Movie Controller
Controller







 PDBj
PDBj

 X-PLOR
X-PLOR