+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-9849 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | LAT1-CD98hc complex bound to MEM-108 Fab | |||||||||
 マップデータ マップデータ | Sharpened, masked map | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | Transporter / Glycoprotein / Complex / MEMBRANE PROTEIN / MEMBRANE PROTEIN-IMMUNE SYSTEM complex | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報L-tryptophan transmembrane transport / positive regulation of L-leucine import across plasma membrane / L-tryptophan transmembrane transporter activity / cellular response to L-arginine / apical pole of neuron / tyrosine transport / L-histidine transport / amino acid transport complex / L-leucine import across plasma membrane / L-alanine transmembrane transporter activity ...L-tryptophan transmembrane transport / positive regulation of L-leucine import across plasma membrane / L-tryptophan transmembrane transporter activity / cellular response to L-arginine / apical pole of neuron / tyrosine transport / L-histidine transport / amino acid transport complex / L-leucine import across plasma membrane / L-alanine transmembrane transporter activity / L-alanine import across plasma membrane / Defective SLC7A7 causes lysinuric protein intolerance (LPI) / aromatic amino acid transmembrane transporter activity / phenylalanine transport / methionine transport / L-leucine transmembrane transporter activity / amino acid transmembrane transport / thyroid hormone transmembrane transporter activity / isoleucine transport / valine transport / alanine transport / proline transport / L-amino acid transmembrane transporter activity / L-leucine transport / thyroid hormone transport / negative regulation of vascular associated smooth muscle cell apoptotic process / neutral amino acid transport / positive regulation of cytokine production involved in immune response / amino acid import across plasma membrane / neutral L-amino acid transmembrane transporter activity / external side of apical plasma membrane / Tryptophan catabolism / Amino acid transport across the plasma membrane / amino acid transmembrane transporter activity / exogenous protein binding / anchoring junction / antiporter activity / Basigin interactions / xenobiotic transport / microvillus membrane / response to muscle activity / positive regulation of interleukin-4 production / response to exogenous dsRNA / amino acid transport / positive regulation of interleukin-17 production / tryptophan transport / positive regulation of glial cell proliferation / response to hyperoxia / transport across blood-brain barrier / cellular response to glucose starvation / liver regeneration / negative regulation of autophagy / basal plasma membrane / peptide antigen binding / positive regulation of type II interferon production / calcium ion transport / melanosome / double-stranded RNA binding / cellular response to lipopolysaccharide / virus receptor activity / basolateral plasma membrane / carbohydrate metabolic process / apical plasma membrane / cadherin binding / protein heterodimerization activity / negative regulation of gene expression / lysosomal membrane / intracellular membrane-bounded organelle / synapse / symbiont entry into host cell / cell surface / protein homodimerization activity / RNA binding / extracellular exosome / nucleoplasm / membrane / plasma membrane / cytosol 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) / Homo sapiens (ヒト) /  | |||||||||
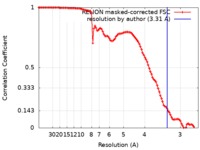
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.31 Å | |||||||||
 データ登録者 データ登録者 | Lee Y / Nishizawa T | |||||||||
| 資金援助 |  日本, 2件 日本, 2件
| |||||||||
 引用 引用 |  ジャーナル: Nat Struct Mol Biol / 年: 2019 ジャーナル: Nat Struct Mol Biol / 年: 2019タイトル: Cryo-EM structure of the human L-type amino acid transporter 1 in complex with glycoprotein CD98hc. 著者: Yongchan Lee / Pattama Wiriyasermkul / Chunhuan Jin / Lili Quan / Ryuichi Ohgaki / Suguru Okuda / Tsukasa Kusakizako / Tomohiro Nishizawa / Kazumasa Oda / Ryuichiro Ishitani / Takeshi ...著者: Yongchan Lee / Pattama Wiriyasermkul / Chunhuan Jin / Lili Quan / Ryuichi Ohgaki / Suguru Okuda / Tsukasa Kusakizako / Tomohiro Nishizawa / Kazumasa Oda / Ryuichiro Ishitani / Takeshi Yokoyama / Takanori Nakane / Mikako Shirouzu / Hitoshi Endou / Shushi Nagamori / Yoshikatsu Kanai / Osamu Nureki /    要旨: The L-type amino acid transporter 1 (LAT1 or SLC7A5) transports large neutral amino acids across the membrane and is crucial for brain drug delivery and tumor growth. LAT1 forms a disulfide-linked ...The L-type amino acid transporter 1 (LAT1 or SLC7A5) transports large neutral amino acids across the membrane and is crucial for brain drug delivery and tumor growth. LAT1 forms a disulfide-linked heterodimer with CD98 heavy chain (CD98hc, 4F2hc or SLC3A2), but the mechanism of assembly and amino acid transport are poorly understood. Here we report the cryo-EM structure of the human LAT1-CD98hc heterodimer at 3.3-Å resolution. LAT1 features a canonical Leu T-fold and exhibits an unusual loop structure on transmembrane helix 6, creating an extended cavity that might accommodate bulky amino acids and drugs. CD98hc engages with LAT1 through the extracellular, transmembrane and putative cholesterol-mediated interactions. We also show that two anti-CD98 antibodies recognize distinct, multiple epitopes on CD98hc but not its glycans, explaining their robust reactivities. These results reveal the principles of glycoprotein-solute carrier assembly and provide templates for improving preclinical drugs and antibodies targeting LAT1 or CD98hc. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_9849.map.gz emd_9849.map.gz | 5.5 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-9849-v30.xml emd-9849-v30.xml emd-9849.xml emd-9849.xml | 29.2 KB 29.2 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_9849_fsc.xml emd_9849_fsc.xml | 10.2 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_9849.png emd_9849.png | 94.1 KB | ||
| マスクデータ |  emd_9849_msk_1.map emd_9849_msk_1.map | 91.1 MB |  マスクマップ マスクマップ | |
| Filedesc metadata |  emd-9849.cif.gz emd-9849.cif.gz | 8.6 KB | ||
| その他 |  emd_9849_half_map_1.map.gz emd_9849_half_map_1.map.gz emd_9849_half_map_2.map.gz emd_9849_half_map_2.map.gz | 71.3 MB 71.2 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9849 http://ftp.pdbj.org/pub/emdb/structures/EMD-9849 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9849 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9849 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_9849_validation.pdf.gz emd_9849_validation.pdf.gz | 165.1 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_9849_full_validation.pdf.gz emd_9849_full_validation.pdf.gz | 164.6 KB | 表示 | |
| XML形式データ |  emd_9849_validation.xml.gz emd_9849_validation.xml.gz | 499 B | 表示 | |
| CIF形式データ |  emd_9849_validation.cif.gz emd_9849_validation.cif.gz | 512 B | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9849 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9849 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9849 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9849 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  6jmqMC  9850C  6jmrC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | |
| 電子顕微鏡画像生データ |  EMPIAR-10264 (タイトル: LAT1-CD98hc bound to MEM-108 Fab / Data size: 4.1 TB EMPIAR-10264 (タイトル: LAT1-CD98hc bound to MEM-108 Fab / Data size: 4.1 TBData #1: Unaligned multi-frame micrographs, Dataset1 [micrographs - multiframe] Data #2: Unaligned multi-frame micrographs, Dataset2 [micrographs - multiframe] Data #3: Micelle subtracted particles used for final reconstruction [picked particles - single frame - processed]) |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_9849.map.gz / 形式: CCP4 / 大きさ: 91.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_9849.map.gz / 形式: CCP4 / 大きさ: 91.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Sharpened, masked map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.333 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-マスク #1
| ファイル |  emd_9849_msk_1.map emd_9849_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
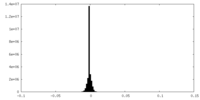
| 密度ヒストグラム |
-ハーフマップ: Half map 2 used for FSC calculation. The...
| ファイル | emd_9849_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Half map 2 used for FSC calculation. The header contains a wrong pixel size information (1.375 A). The correct pixel size is 1.333 A. | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
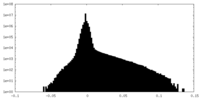
| 密度ヒストグラム |
-ハーフマップ: Half map 1 used for FSC calculation. The...
| ファイル | emd_9849_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Half map 1 used for FSC calculation. The header contains a wrong pixel size information (1.375 A). The correct pixel size is 1.333 A. | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : LAT1-CD98hc complex bound to MEM-108 Fab
| 全体 | 名称: LAT1-CD98hc complex bound to MEM-108 Fab |
|---|---|
| 要素 |
|
-超分子 #1: LAT1-CD98hc complex bound to MEM-108 Fab
| 超分子 | 名称: LAT1-CD98hc complex bound to MEM-108 Fab / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1-#4 |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 175 kDa/nm |
-分子 #1: Large neutral amino acids transporter small subunit 1
| 分子 | 名称: Large neutral amino acids transporter small subunit 1 タイプ: protein_or_peptide / ID: 1 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 56.043746 KDa |
| 組換発現 | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 配列 | 文字列: MAGAGPKRRA LAAPAAEEKE EAREKMLAAK SADGSAPAGE GEGVTLQRNI TLLNGVAIIV GTIIGSGIFV TPTGVLKEAG SPGLALVVW AACGVFSIVG ALCYAELGTT ISKSGGDYAY MLEVYGSLPA FLKLWIELLI IRPSSQYIVA LVFATYLLKP L FPTCPVPE ...文字列: MAGAGPKRRA LAAPAAEEKE EAREKMLAAK SADGSAPAGE GEGVTLQRNI TLLNGVAIIV GTIIGSGIFV TPTGVLKEAG SPGLALVVW AACGVFSIVG ALCYAELGTT ISKSGGDYAY MLEVYGSLPA FLKLWIELLI IRPSSQYIVA LVFATYLLKP L FPTCPVPE EAAKLVACLC VLLLTAVNCY SVKAATRVQD AFAAAKLLAL ALIILLGFVQ IGKGDVSNLD PNFSFEGTKL DV GNIVLAL YSGLFAYGGW NYLNFVTEEM INPYRNLPLA IIISLPIVTL VYVLTNLAYF TTLSTEQMLS SEAVAVDFGN YHL GVMSWI IPVFVGLSCF GSVNGSLFTS SRLFFVGSRE GHLPSILSMI HPQLLTPVPS LVFTCVMTLL YAFSKDIFSV INFF SFFNW LCVALAIIGM IWLRHRKPEL ERPIKVNLAL PVFFILACLF LIAVSFWKTP VECGIGFTII LSGLPVYFFG VWWKN KPKW LLQGIFSTTV LCQKLMQVVP QETDYKDDDD K UniProtKB: Large neutral amino acids transporter small subunit 1 |
-分子 #2: 4F2 cell-surface antigen heavy chain
| 分子 | 名称: 4F2 cell-surface antigen heavy chain / タイプ: protein_or_peptide / ID: 2 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 68.069625 KDa |
| 組換発現 | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 配列 | 文字列: GSELQPPEAS IAVVSIPRQL PGSHSEAGVQ GLSAGDDSEL GSHCVAQTGL ELLASGDPLP SASQNAEMIE TGSDCVTQAG LQLLASSDP PALASKNAEV TGTMSQDTEV DMKEVELNEL EPEKQPMNAA SGAAMSLAGA EKNGLVKIKV AEDEAEAAAA A KFTGLSKE ...文字列: GSELQPPEAS IAVVSIPRQL PGSHSEAGVQ GLSAGDDSEL GSHCVAQTGL ELLASGDPLP SASQNAEMIE TGSDCVTQAG LQLLASSDP PALASKNAEV TGTMSQDTEV DMKEVELNEL EPEKQPMNAA SGAAMSLAGA EKNGLVKIKV AEDEAEAAAA A KFTGLSKE ELLKVAGSPG WVRTRWALLL LFWLGWLGML AGAVVIIVRA PRCRELPAQK WWHTGALYRI GDLQAFQGHG AG NLAGLKG RLDYLSSLKV KGLVLGPIHK NQKDDVAQTD LLQIDPNFGS KEDFDSLLQS AKKKSIRVIL DLTPNYRGEN SWF STQVDT VATKVKDALE FWLQAGVDGF QVRDIENLKD ASSFLAEWQN ITKGFSEDRL LIAGTNSSDL QQILSLLESN KDLL LTSSY LSDSGSTGEH TKSLVTQYLN ATGNRWCSWS LSQARLLTSF LPAQLLRLYQ LMLFTLPGTP VFSYGDEIGL DAAAL PGQP MEAPVMLWDE SSFPDIPGAV SANMTVKGQS EDPGSLLSLF RRLSDQRSKE RSLLHGDFHA FSAGPGLFSY IRHWDQ NER FLVVLNFGDV GLSAGLQASD LPASASLPAK ADLLLSTQPG REEGSPLELE RLKLEPHEGL LLRFPYAA UniProtKB: Amino acid transporter heavy chain SLC3A2 |
-分子 #3: Antibody
| 分子 | 名称: Antibody / タイプ: protein_or_peptide / ID: 3 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 22.577857 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: QVQLKESGPG LVAPSQSLSI TCTVSGFPLT (UNK)(UNK)(UNK)(UNK)(UNK)WVRQP PGKGLEWLG(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)RLSI SKDNSK SQV FLKMNSLQTD ...文字列: QVQLKESGPG LVAPSQSLSI TCTVSGFPLT (UNK)(UNK)(UNK)(UNK)(UNK)WVRQP PGKGLEWLG(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)RLSI SKDNSK SQV FLKMNSLQTD DTARYYCAR(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)W GQGTSVT VS SAKTTPPSVY PLAPGS(UNK)(UNK)(UNK)(UNK) (UNK)SMVTLGCLV KGYFPEPVTV TWNSGSLSSG VHTFPAVL Q SDLYTLSSSV TVPSSTWPSE TVTCNVAHPA SSTKVDKKIV PRD |
-分子 #4: Antibody
| 分子 | 名称: Antibody / タイプ: protein_or_peptide / ID: 4 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 23.227152 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: DIVMSQSPSS LVVSVGEKVT MSC(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)WYQQKPGQS PKLLIY(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) GVPDRF TGSGSGTDFT ...文字列: DIVMSQSPSS LVVSVGEKVT MSC(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)WYQQKPGQS PKLLIY(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) GVPDRF TGSGSGTDFT LTISSVKAED LAVYYC(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)FGG G TKLEIKRADA APTVSIFPPS SEQLTSGGAS VVCFLNNFYP KDINVKWKID GSERQNGVLN SWTDQDSKDS TYSMSSTLT LTKDEYERHN SYTCEATHKT STSPIVKSFN RNE |
-分子 #6: CHOLESTEROL
| 分子 | 名称: CHOLESTEROL / タイプ: ligand / ID: 6 / コピー数: 5 / 式: CLR |
|---|---|
| 分子量 | 理論値: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-分子 #7: 2-acetamido-2-deoxy-beta-D-glucopyranose
| 分子 | 名称: 2-acetamido-2-deoxy-beta-D-glucopyranose / タイプ: ligand / ID: 7 / コピー数: 3 / 式: NAG |
|---|---|
| 分子量 | 理論値: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 8 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: FEI FALCON III (4k x 4k) 平均電子線量: 7.14 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)