[English] 日本語
 Yorodumi
Yorodumi- EMDB-9742: Cryo-EM structure of AcrVA5-acetylated MbCas12a in complex with crRNA -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9742 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of AcrVA5-acetylated MbCas12a in complex with crRNA | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | enzyme / IMMUNE SYSTEM / IMMUNE SYSTEM-RNA complex | |||||||||
| Function / homology | CRISPR-associated endonuclease Cas12a / Cas12a, REC1 domain / Cas12a, RuvC nuclease domain / Cas12a, nuclease domain / Alpha helical recognition lobe domain / Nuclease domain / RuvC nuclease domain / Type V CRISPR-associated protein Cpf1 Function and homology information Function and homology information | |||||||||
| Biological species |  Moraxella bovoculi (bacteria) / synthetic construct (others) Moraxella bovoculi (bacteria) / synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Dong L / Li N | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2019 Journal: Nat Struct Mol Biol / Year: 2019Title: An anti-CRISPR protein disables type V Cas12a by acetylation. Authors: Liyong Dong / Xiaoyu Guan / Ningning Li / Fan Zhang / Yuwei Zhu / Kuan Ren / Ling Yu / Fengxia Zhou / Zhifu Han / Ning Gao / Zhiwei Huang /  Abstract: Phages use anti-CRISPR proteins to deactivate the CRISPR-Cas system. The mechanisms for the inhibition of type I and type II systems by anti-CRISPRs have been elucidated. However, it has remained ...Phages use anti-CRISPR proteins to deactivate the CRISPR-Cas system. The mechanisms for the inhibition of type I and type II systems by anti-CRISPRs have been elucidated. However, it has remained unknown how the type V CRISPR-Cas12a (Cpf1) system is inhibited by anti-CRISPRs. Here we identify the anti-CRISPR protein AcrVA5 and report the mechanisms by which it inhibits CRISPR-Cas12a. Our structural and biochemical data show that AcrVA5 functions as an acetyltransferase to modify Moraxella bovoculi (Mb) Cas12a at Lys635, a residue that is required for recognition of the protospacer-adjacent motif. The AcrVA5-mediated modification of MbCas12a results in complete loss of double-stranded DNA (dsDNA)-cleavage activity. In contrast, the Lys635Arg mutation renders MbCas12a completely insensitive to inhibition by AcrVA5. A cryo-EM structure of the AcrVA5-acetylated MbCas12a reveals that Lys635 acetylation provides sufficient steric hindrance to prevent dsDNA substrates from binding to the Cas protein. Our study reveals an unprecedented mechanism of CRISPR-Cas inhibition and suggests an evolutionary arms race between phages and bacteria. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9742.map.gz emd_9742.map.gz | 5.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9742-v30.xml emd-9742-v30.xml emd-9742.xml emd-9742.xml | 15.1 KB 15.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9742.png emd_9742.png | 90.4 KB | ||
| Filedesc metadata |  emd-9742.cif.gz emd-9742.cif.gz | 7.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9742 http://ftp.pdbj.org/pub/emdb/structures/EMD-9742 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9742 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9742 | HTTPS FTP |
-Related structure data
| Related structure data |  6iv6MC  6iufC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_9742.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9742.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
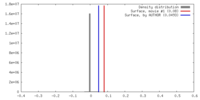
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : cpf1-crRNA
| Entire | Name: cpf1-crRNA |
|---|---|
| Components |
|
-Supramolecule #1: cpf1-crRNA
| Supramolecule | Name: cpf1-crRNA / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Moraxella bovoculi (bacteria) Moraxella bovoculi (bacteria) |
-Macromolecule #1: nuclease
| Macromolecule | Name: nuclease / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Moraxella bovoculi (bacteria) Moraxella bovoculi (bacteria) |
| Molecular weight | Theoretical: 145.253391 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MLFQDFTHLY PLSKTVRFEL KPIG(ALY)TLEHI HAKNFLNQDE TMADMYQKVK AILDDYHRDF IADMMGEVKL TKLAEF YDV YLKFRKNPKD DGLQKQLKDL QAVLRKEIVK PIGNGGKYKA GYDRLFGAKL FKDGKELGDL AKFVIAQEGE SSP (ALY)LAHLA ...String: MLFQDFTHLY PLSKTVRFEL KPIG(ALY)TLEHI HAKNFLNQDE TMADMYQKVK AILDDYHRDF IADMMGEVKL TKLAEF YDV YLKFRKNPKD DGLQKQLKDL QAVLRKEIVK PIGNGGKYKA GYDRLFGAKL FKDGKELGDL AKFVIAQEGE SSP (ALY)LAHLA HFEKFSTYFT GFHDNRKNMY SDEDKHTAIA YRLIHENLPR FIDNLQILAT IKQKHSALYD QIINELTASG LDVSLASHL DGYHKLLTQE GITAYNTLLG GISGEAGSRK IQGINELINS HHNQHCHKSE RIAKLRPLHK QILSDGMGVS F LPSKFADD SEVCQAVNEF YRHYADVFAK VQSLFDGFDD YQKDGIYVEY KNLNELSKQA FGDFALLGRV LDGYYVDVVN PE FNERFAK AKTDNAKAKL TKEKDKFIKG VHSLASLEQA IEHYTARHDD ESVQAGKLGQ YFKHGLAGVD NPIQKIHNNH STI KGFLER ERPAGERALP KIKSDKSPEI RQLKELLDNA LNVAHFAKLL TTKTTLHNQD GNFYGEFGAL YDELAKIATL YNKV RDYLS QKPFSTEKYK LNFGNPTLLN GWDLNKEKDN FGVILQKDGC YYLALLDKAH KKVFDNAPNT GKSVYQKMIY KLLPG PNKM LP(ALY)VFFAKSN LDYYNPSAEL LDKYAQGTHK KGDNFNLKDC HALIDFFKAG INKHPEWQHF GFKFSPTSSY QD LSDFYRE VEPQGYQVKF VDINADYINE LVEQGQLYLF QIYNKDFSPK AHGKPNLHTL YFKALFSEDN LVNPIYKLNG EAE IFYRKA SLDMNETTIH RAGEVLENKN PDNPK(ALY)RQFV YDIIKDKRYT QDKFMLHVPI TMNFGVQGMT IKEFNKKVNQ SIQQYDEVN VIGIDRGERH LLYLTVINSK GEILEQRSLN DITTASANGT QMTTPYHKIL DKREIERLNA RVGWGEIETI K ELKSGYLS HVVHQISQLM LKYNAIVVLE DLNFGFKRGC FKVEKQIYQN FENALIKKLN HLVLKDKADD EIGSYKNALQ LT NNFTDLK SIGKQTGFLF YVPAWNTSKI DPETGFVDLL KPRYENIAQS QAFFGKFDKI CYNADRGYFE FHIDYAKFND KAK NSRQIW KICSHGDKRY VYDKTANQNK GATIGVNVND ELKSLFTRYH INDKQPNLVM DICQNNDKEF HKSLMYLLKT LLAL RYSNA SSDEDFILSP VANDEGVFFN SALADDTQPQ NADANGAYHI ALKGLWLLNE LKNSDDLNKV KLAIDNQTWL NFAQN R UniProtKB: Type V CRISPR-associated protein Cpf1 |
-Macromolecule #2: RNA (59-MER)
| Macromolecule | Name: RNA (59-MER) / type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 18.673896 KDa |
| Sequence | String: GUCUAACGAC CUUUUAAAUU UCUACUGUUU GUAGAUCUGA UGGUCCAUGU CUGUUACUC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: NITROGEN |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 81.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)