+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7999 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Ion channel receptor | |||||||||
 Map data Map data | Ion channel receptor | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationTRP channels / Neutrophil degranulation / ADP-D-ribose binding / mono-ADP-D-ribose binding / ligand-gated calcium channel activity / ligand-gated monoatomic cation channel activity / monoatomic ion channel activity / calcium ion transmembrane transport / calcium channel activity / protein homotetramerization ...TRP channels / Neutrophil degranulation / ADP-D-ribose binding / mono-ADP-D-ribose binding / ligand-gated calcium channel activity / ligand-gated monoatomic cation channel activity / monoatomic ion channel activity / calcium ion transmembrane transport / calcium channel activity / protein homotetramerization / calcium ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Du J / Lu W | |||||||||
 Citation Citation |  Journal: Nature / Year: 2018 Journal: Nature / Year: 2018Title: Architecture of the TRPM2 channel and its activation mechanism by ADP-ribose and calcium. Authors: Yihe Huang / Paige A Winkler / Weinan Sun / Wei Lü / Juan Du /  Abstract: Transient receptor potential melastatin 2 (TRPM2) is a calcium-permeable, non-selective cation channel that has an essential role in diverse physiological processes such as core body temperature ...Transient receptor potential melastatin 2 (TRPM2) is a calcium-permeable, non-selective cation channel that has an essential role in diverse physiological processes such as core body temperature regulation, immune response and apoptosis. TRPM2 is polymodal and can be activated by a wide range of stimuli, including temperature, oxidative stress and NAD-related metabolites such as ADP-ribose (ADPR). Its activation results in both Ca entry across the plasma membrane and Ca release from lysosomes, and has been linked to diseases such as ischaemia-reperfusion injury, bipolar disorder and Alzheimer's disease. Here we report the cryo-electron microscopy structures of the zebrafish TRPM2 in the apo resting (closed) state and in the ADPR/Ca-bound active (open) state, in which the characteristic NUDT9-H domains hang underneath the MHR1/2 domain. We identify an ADPR-binding site located in the bi-lobed structure of the MHR1/2 domain. Our results provide an insight into the mechanism of activation of the TRPM channel family and define a framework for the development of therapeutic agents to treat neurodegenerative diseases and temperature-related pathological conditions. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7999.map.gz emd_7999.map.gz | 95.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7999-v30.xml emd-7999-v30.xml emd-7999.xml emd-7999.xml | 14.8 KB 14.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7999.png emd_7999.png | 312.8 KB | ||
| Filedesc metadata |  emd-7999.cif.gz emd-7999.cif.gz | 6.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7999 http://ftp.pdbj.org/pub/emdb/structures/EMD-7999 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7999 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7999 | HTTPS FTP |
-Related structure data
| Related structure data |  6drjMC  8901C  6drkC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_7999.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7999.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Ion channel receptor | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.074 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Ion channel 1
| Entire | Name: Ion channel 1 |
|---|---|
| Components |
|
-Supramolecule #1: Ion channel 1
| Supramolecule | Name: Ion channel 1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 500 KDa |
-Macromolecule #1: Transient receptor potential cation channel, subfamily M, member 2
| Macromolecule | Name: Transient receptor potential cation channel, subfamily M, member 2 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 168.601672 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MLGTSGVKIH PNGNSNQLGV QLENVKLTSL FKKLDKRCSL ASWIKENIKK KECCFYVEDG REGICKCGYP KVQHCDEAIK PEDYMGEQW DKHRHVRETP TDAFGDISFG GLGQKTGKYV RVSSDTSCEN LYQLMTEQWK LRSPNLLISV TGGAKNFYIK T HLKDKFRR ...String: MLGTSGVKIH PNGNSNQLGV QLENVKLTSL FKKLDKRCSL ASWIKENIKK KECCFYVEDG REGICKCGYP KVQHCDEAIK PEDYMGEQW DKHRHVRETP TDAFGDISFG GLGQKTGKYV RVSSDTSCEN LYQLMTEQWK LRSPNLLISV TGGAKNFYIK T HLKDKFRR GLIKVAQTTG AWILTGGTHA GVMKHVGMAV RDYTLSSGSM EGQIVVIGVA PWGVIHNRST LIHPEGRFPA YY SLDEQGQ GRLSCLDINH THFLLVDDGT QGHYGVEIEL RARLEKLISK LSLGNRESGV TIPVVCVVLD GGPGTLNTIY NSM LNHTPC VVLEGSGRLA DVIAHVASVP VSKVTMALIN RLLKRFFMQE YKNFTELQII EWTKKIQDIL RMPHLLTVFR IDED KNYDV DVAILQALLK ASRSDEHAGR HCWERQLELA VAWNRVDIAE SEIFTEESQW TSSDLHPAMF SALVGDKPEF VRLLL ENGV CVREFLEREE TLCELYSHLP SCFFLRKLAK RVQGGKMRRG QEPLPGSRKV CLSHVSEEVR HLLGSFTQPL YIASRY KPT KDDVRLKVPS KGALDLPCSG EEWSADTVWD PGRDLFLWAV VQNNRELAEI GWEQCRDCIA AALAASKILR KLAQESG ED DSEEATEMLE LANHYEKQAI GVFSECHSWD AQRAQKLLIR ISPSWGRSTC LWLALEAHDK SFIAHSGVQA LLTQIWCG E LSVDNPHWKV LLCMIFFPLI YTGFLTFRRD EDIQRQAERT QQKLAMESVF AGQSDGKIKR HLRGFSQKSE LKPLNCSSR LMSFLKSPQV KFYWNIASYF GFLWLFAVVL MIDFQTSPSW RELLLYVWLT SLVCEEIRQL YHDFDGSGFR RKAKMYIKDL WNILDVLSI VLFIAGLICR LQASDTVFYI GKVILCIDFI IFCLRLMAIF SISRTLGPKI IIVRRMMLDL FFFMFLLSIW V VAYGVAKQ GILIENEERL NWIIRGAVYE PYITIFGNFP TNIDNTLFDI SSCSVNASDP LKPKCPMLNA DNTPVFPEWL TI MMLCVYL LFANILLLNL LIAIFNYTFQ EVQDNTDTIW KFQRYELIKE YHSRPALPPP FILLSHLILF IRGVFLRDLP QRH KNFRQE LEQTEEEELL SWEAYMKDNY LASTRQDESQ SVEHRIHDTA EKVGAMSELL EREQEMVSAT MAKRLARLEE QVSE SAKAL RWIIDALKSQ GCKSKVQPPL MRSKSSDRDD GDSSGQETDD EEAPHMFARQ LQYPDSTVRR FPVPEEKVSW EVNFS PYQP PVYNQQDSSE SDTSALDKHR NPGGRTGIRG KGALNTLGPN HILHPIFTRW RDAEHKVLEF LAVWEDAEKR WALLGG PAQ PDEPLAQVLE RILGKKLNEK TKTLLKAGEE VYKGYVDDSR NTDNAWVETS IITLHCDKNT PLMADLNHMV ESSLSSH QP LQWREVSSDA CRCSYQREAL RQIAHHHNTY F UniProtKB: UNIPROTKB: A0A0R4IN04 |
-Macromolecule #2: ADENOSINE-5-DIPHOSPHORIBOSE
| Macromolecule | Name: ADENOSINE-5-DIPHOSPHORIBOSE / type: ligand / ID: 2 / Number of copies: 4 / Formula: APR |
|---|---|
| Molecular weight | Theoretical: 559.316 Da |
| Chemical component information | 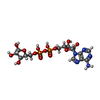 ChemComp-APR: |
-Macromolecule #3: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 3 / Number of copies: 4 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















