[English] 日本語
 Yorodumi
Yorodumi- EMDB-7788: Insulin Receptor ectodomain in complex with two insulin molecules... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7788 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Insulin Receptor ectodomain in complex with two insulin molecules plunged with a spot-to-plunge time of 200 ms | |||||||||
 Map data Map data | CryoSPARC auto-generated sharpened full map from the last iteration of independent refinement. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.93 Å | |||||||||
 Authors Authors | Noble AJ / Wei H / Dandey VP / Zhang Z / Potter CS / Carragher B | |||||||||
 Citation Citation |  Journal: Nat Methods / Year: 2018 Journal: Nat Methods / Year: 2018Title: Reducing effects of particle adsorption to the air-water interface in cryo-EM. Authors: Alex J Noble / Hui Wei / Venkata P Dandey / Zhening Zhang / Yong Zi Tan / Clinton S Potter / Bridget Carragher /  Abstract: Most protein particles prepared in vitreous ice for single-particle cryo-electron microscopy (cryo-EM) are adsorbed to air-water or substrate-water interfaces, which can cause the particles to adopt ...Most protein particles prepared in vitreous ice for single-particle cryo-electron microscopy (cryo-EM) are adsorbed to air-water or substrate-water interfaces, which can cause the particles to adopt preferred orientations. By using a rapid plunge-freezing robot and nanowire grids, we were able to reduce some of the deleterious effects of the air-water interface by decreasing the dwell time of particles in thin liquid films. We demonstrated this by using single-particle cryo-EM and cryo-electron tomography (cryo-ET) to examine hemagglutinin, insulin receptor complex, and apoferritin. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7788.map.gz emd_7788.map.gz | 86 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7788-v30.xml emd-7788-v30.xml emd-7788.xml emd-7788.xml | 15.8 KB 15.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7788.png emd_7788.png | 62.4 KB | ||
| Masks |  emd_7788_msk_1.map emd_7788_msk_1.map | 91.1 MB |  Mask map Mask map | |
| Others |  emd_7788_half_map_1.map.gz emd_7788_half_map_1.map.gz emd_7788_half_map_2.map.gz emd_7788_half_map_2.map.gz | 84.5 MB 84.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7788 http://ftp.pdbj.org/pub/emdb/structures/EMD-7788 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7788 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7788 | HTTPS FTP |
-Related structure data
| Related structure data |  7623C  7624C  7625C  7627C  7628C  7629C  7630C  7791C  7792C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_7788.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7788.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoSPARC auto-generated sharpened full map from the last iteration of independent refinement. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
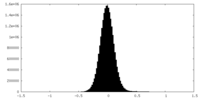
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_7788_msk_1.map emd_7788_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
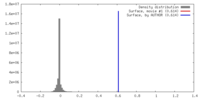
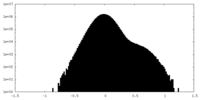
| Density Histograms |
-Half map: CryoSPARC auto-generated half-map from the last iteration of...
| File | emd_7788_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoSPARC auto-generated half-map from the last iteration of independent refinement. | ||||||||||||
| Projections & Slices |
| ||||||||||||
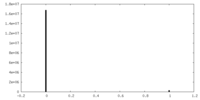
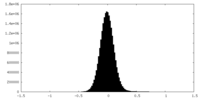
| Density Histograms |
-Half map: CryoSPARC auto-generated half-map from the last iteration of...
| File | emd_7788_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoSPARC auto-generated half-map from the last iteration of independent refinement. | ||||||||||||
| Projections & Slices |
| ||||||||||||
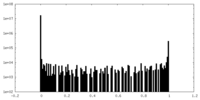
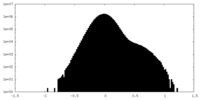
| Density Histograms |
- Sample components
Sample components
-Entire : Insulin Receptor Ectodomain in complex with two insulin molecules
| Entire | Name: Insulin Receptor Ectodomain in complex with two insulin molecules |
|---|---|
| Components |
|
-Supramolecule #1: Insulin Receptor Ectodomain in complex with two insulin molecules
| Supramolecule | Name: Insulin Receptor Ectodomain in complex with two insulin molecules type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Molecular weight | Experimental: 215 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Homemade |
| Vitrification | Cryogen name: ETHANE / Instrument: OTHER Details: 200 ms spot-to-plunge time. The value given for _emd_vitrification.instrument is SPOTITON. This is not in a list of allowed values set(['LEICA EM CPC', 'GATAN CRYOPLUNGE 3', 'LEICA PLUNGER', ...Details: 200 ms spot-to-plunge time. The value given for _emd_vitrification.instrument is SPOTITON. This is not in a list of allowed values set(['LEICA EM CPC', 'GATAN CRYOPLUNGE 3', 'LEICA PLUNGER', 'FEI VITROBOT MARK II', 'HOMEMADE PLUNGER', 'REICHERT-JUNG PLUNGER', 'FEI VITROBOT MARK I', 'LEICA KF80', 'FEI VITROBOT MARK III', 'LEICA EM GP', 'OTHER', 'FEI VITROBOT MARK IV']) so OTHER is written into the XML file. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Spherical aberration corrector: Corrected so that Cs is as close to 0 as possible. Energy filter - Name: GIF Bioquantum |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Average exposure time: 10.0 sec. / Average electron dose: 66.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 0.01 mm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)