[English] 日本語
 Yorodumi
Yorodumi- EMDB-7515: Subtomogram averages of budding-arrested nucleocapsid-like partic... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7515 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Subtomogram averages of budding-arrested nucleocapsid-like particles (NCLPs) inside CHIKV-infected human cells. | |||||||||
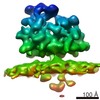
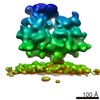
 Map data Map data | Subtomogram average of budding-arrested nucleocapsid-like particles (NCLPs) inside CHIKV-infected human cells. | |||||||||
 Sample Sample | Budding-arrestedChikungunya virus nucleocapsid-like particle != Chikungunya virus Budding-arrestedChikungunya virus nucleocapsid-like particle
| |||||||||
| Biological species |   Chikungunya virus Chikungunya virus | |||||||||
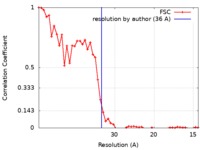
| Method | subtomogram averaging / cryo EM / Resolution: 36.0 Å | |||||||||
 Authors Authors | Galaz-Montoya JG / Jin J / Sherman MB | |||||||||
 Citation Citation |  Journal: Cell Host Microbe / Year: 2018 Journal: Cell Host Microbe / Year: 2018Title: Neutralizing Antibodies Inhibit Chikungunya Virus Budding at the Plasma Membrane. Authors: Jing Jin / Jesús G Galaz-Montoya / Michael B Sherman / Stella Y Sun / Cynthia S Goldsmith / Eileen T O'Toole / Larry Ackerman / Lars-Anders Carlson / Scott C Weaver / Wah Chiu / Graham Simmons /   Abstract: Neutralizing antibodies (NAbs) are traditionally thought to inhibit virus infection by preventing virion entry into target cells. In addition, antibodies can engage Fc receptors (FcRs) on immune ...Neutralizing antibodies (NAbs) are traditionally thought to inhibit virus infection by preventing virion entry into target cells. In addition, antibodies can engage Fc receptors (FcRs) on immune cells to activate antiviral responses. We describe a mechanism by which NAbs inhibit chikungunya virus (CHIKV), the most common alphavirus infecting humans, by preventing virus budding from infected human cells and activating IgG-specific Fcγ receptors. NAbs bind to CHIKV glycoproteins on the infected cell surface and induce glycoprotein coalescence, preventing budding of nascent virions and leaving structurally heterogeneous nucleocapsids arrested in the cytosol. Furthermore, NAbs induce clustering of CHIKV replication spherules at sites of budding blockage. Functionally, these densely packed glycoprotein-NAb complexes on infected cells activate Fcγ receptors, inducing a strong, antibody-dependent, cell-mediated cytotoxicity response from immune effector cells. Our findings describe a triply functional antiviral pathway for NAbs that might be broadly applicable across virus-host systems, suggesting avenues for therapeutic innovation through antibody design. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7515.map.gz emd_7515.map.gz | 599.3 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7515-v30.xml emd-7515-v30.xml emd-7515.xml emd-7515.xml | 19.8 KB 19.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_7515_fsc.xml emd_7515_fsc.xml | 4.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_7515_1.png emd_7515_1.png emd_7515_2.png emd_7515_2.png emd_7515_3.png emd_7515_3.png emd_7515_4.png emd_7515_4.png emd_7515_5.png emd_7515_5.png | 57.6 KB 58.4 KB 65.2 KB 67.4 KB 65.8 KB | ||
| Others |  emd_7515_additional_1.map.gz emd_7515_additional_1.map.gz emd_7515_additional_2.map.gz emd_7515_additional_2.map.gz emd_7515_additional_3.map.gz emd_7515_additional_3.map.gz emd_7515_additional_4.map.gz emd_7515_additional_4.map.gz | 615.5 KB 629.1 KB 664 KB 681.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7515 http://ftp.pdbj.org/pub/emdb/structures/EMD-7515 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7515 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7515 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_7515.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7515.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Subtomogram average of budding-arrested nucleocapsid-like particles (NCLPs) inside CHIKV-infected human cells. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 7.16 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Subtomogram average of budding-arrested nucleocapsid-like particles (NCLPs) inside...
| File | emd_7515_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Subtomogram average of budding-arrested nucleocapsid-like particles (NCLPs) inside CHIKV-infected human cells. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Subtomogram average of budding-arrested nucleocapsid-like particles (NCLPs) inside...
| File | emd_7515_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Subtomogram average of budding-arrested nucleocapsid-like particles (NCLPs) inside CHIKV-infected human cells. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Subtomogram average of budding-arrested nucleocapsid-like particles (NCLPs) inside...
| File | emd_7515_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Subtomogram average of budding-arrested nucleocapsid-like particles (NCLPs) inside CHIKV-infected human cells. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Subtomogram average of budding-arrested nucleocapsid-like particles (NCLPs) inside...
| File | emd_7515_additional_4.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Subtomogram average of budding-arrested nucleocapsid-like particles (NCLPs) inside CHIKV-infected human cells. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Budding-arrestedChikungunya virus nucleocapsid-like particle
| Entire | Name: Budding-arrestedChikungunya virus nucleocapsid-like particle |
|---|---|
| Components |
|
-Supramolecule #1: Chikungunya virus
| Supramolecule | Name: Chikungunya virus / type: virus / ID: 1 / Parent: 0 / NCBI-ID: 37124 / Sci species name: Chikungunya virus / Sci species strain: 181/clone 25 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Virus shell | Shell ID: 1 / Diameter: 698.0 Å / T number (triangulation number): 4 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
| Details | Neutralizing antibody treated U2OS cells that were infected with CHIKV vaccine strain 181/clone 25 |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2200FS |
|---|---|
| Specialist optics | Energy filter - Name: In-column Omega Filter / Energy filter - Lower energy threshold: 0 eV / Energy filter - Upper energy threshold: 20 eV |
| Image recording | Film or detector model: DIRECT ELECTRON DE-20 (5k x 3k) / Average exposure time: 0.6 sec. / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)