+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7455 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the cargo bound AP-1:Arf1:tetherin-Nef monomer | |||||||||
 Map data Map data | Structure of the cargo bound AP-1:Arf1:tetherin-Nef monomer | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of plasmacytoid dendritic cell cytokine production / negative regulation of intracellular transport of viral material / response to interferon-beta / basolateral protein secretion / synaptic vesicle budding from endosome / Lysosome Vesicle Biogenesis / positive regulation of natural killer cell degranulation / AP-1 adaptor complex / endosome to melanosome transport / mitotic cleavage furrow ingression ...negative regulation of plasmacytoid dendritic cell cytokine production / negative regulation of intracellular transport of viral material / response to interferon-beta / basolateral protein secretion / synaptic vesicle budding from endosome / Lysosome Vesicle Biogenesis / positive regulation of natural killer cell degranulation / AP-1 adaptor complex / endosome to melanosome transport / mitotic cleavage furrow ingression / trans-Golgi Network Vesicle Budding / response to interferon-alpha / protein trimerization / platelet dense granule organization / metalloendopeptidase inhibitor activity / negative regulation of glycoprotein biosynthetic process / symbiont-mediated suppression of host antigen processing and presentation of peptide antigen via MHC class I / melanosome assembly / Glycosphingolipid transport / regulation of receptor internalization / symbiont-mediated suppression of host antigen processing and presentation of peptide antigen via MHC class II / Golgi to lysosome transport / Intra-Golgi traffic / Golgi to vacuole transport / regulation of Arp2/3 complex-mediated actin nucleation / Golgi Associated Vesicle Biogenesis / Synthesis of PIPs at the Golgi membrane / symbiont-mediated suppression of host autophagy / symbiont-mediated suppression of host apoptosis / clathrin-cargo adaptor activity / positive regulation of leukocyte proliferation / melanosome organization / GTP-dependent protein binding / MHC class II antigen presentation / thioesterase binding / Nef Mediated CD4 Down-regulation / positive regulation of natural killer cell mediated cytotoxicity / CD4 receptor binding / dendritic spine organization / determination of left/right symmetry / clathrin-coated vesicle / long-term synaptic depression / COPI-dependent Golgi-to-ER retrograde traffic / azurophil granule membrane / negative regulation of viral genome replication / Lysosome Vesicle Biogenesis / clathrin binding / Golgi Associated Vesicle Biogenesis / Synthesis of PIPs at the plasma membrane / cell leading edge / B cell activation / host cell Golgi membrane / MHC class I protein binding / intracellular copper ion homeostasis / response to type II interferon / kinesin binding / synaptic vesicle endocytosis / protein targeting / side of membrane / COPI-mediated anterograde transport / collagen binding / viral life cycle / vesicle-mediated transport / multivesicular body / regulation of calcium-mediated signaling / Neutrophil degranulation / clathrin-coated pit / MHC class II antigen presentation / Gene and protein expression by JAK-STAT signaling after Interleukin-12 stimulation / cytoplasmic vesicle membrane / Nef mediated downregulation of MHC class I complex cell surface expression / negative regulation of cell migration / trans-Golgi network membrane / small monomeric GTPase / sarcomere / regulation of actin cytoskeleton organization / kidney development / trans-Golgi network / intracellular protein transport / negative regulation of cell growth / cellular response to virus / SH3 domain binding / recycling endosome / virion component / small GTPase binding / response to virus / SARS-CoV-1 activates/modulates innate immune responses / Interferon alpha/beta signaling / synaptic vesicle / heart development / ATPase binding / presynapse / G protein activity / defense response to virus / early endosome / positive regulation of canonical NF-kappaB signal transduction / apical plasma membrane / neuron projection / postsynaptic density / symbiont-mediated suppression of host innate immune response Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | |||||||||
 Authors Authors | Morris KL / Buffalo CZ / Hurley JH | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2018 Journal: Cell / Year: 2018Title: HIV-1 Nefs Are Cargo-Sensitive AP-1 Trimerization Switches in Tetherin Downregulation. Authors: Kyle L Morris / Cosmo Z Buffalo / Christina M Stürzel / Elena Heusinger / Frank Kirchhoff / Xuefeng Ren / James H Hurley /   Abstract: The HIV accessory protein Nef counteracts immune defenses by subverting coated vesicle pathways. The 3.7 Å cryo-EM structure of a closed trimer of the clathrin adaptor AP-1, the small GTPase Arf1, ...The HIV accessory protein Nef counteracts immune defenses by subverting coated vesicle pathways. The 3.7 Å cryo-EM structure of a closed trimer of the clathrin adaptor AP-1, the small GTPase Arf1, HIV-1 Nef, and the cytosolic tail of the restriction factor tetherin suggested a mechanism for inactivating tetherin by Golgi retention. The 4.3 Å structure of a mutant Nef-induced dimer of AP-1 showed how the closed trimer is regulated by the dileucine loop of Nef. HDX-MS and mutational analysis were used to show how cargo dynamics leads to alternative Arf1 trimerization, directing Nef targets to be either retained at the trans-Golgi or sorted to lysosomes. Phosphorylation of the NL4-3 M-Nef was shown to regulate AP-1 trimerization, explaining how O-Nefs lacking this phosphosite counteract tetherin but most M-Nefs do not. These observations show how the higher-order organization of a vesicular coat can be allosterically modulated to direct cargoes to distinct fates. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7455.map.gz emd_7455.map.gz | 201.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7455-v30.xml emd-7455-v30.xml emd-7455.xml emd-7455.xml | 12 KB 12 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7455.png emd_7455.png | 644.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7455 http://ftp.pdbj.org/pub/emdb/structures/EMD-7455 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7455 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7455 | HTTPS FTP |
-Related structure data
| Related structure data |  6dffMC  7453C  7454C  7456C  7457C  7458C  7563C  6cm9C  6criC  6d83C  6d84C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10176 (Title: Single particle cryo-EM dataset of the flexible and variable oligomeric state complex AP-1:Arf1:tetherin-HIV-Nef EMPIAR-10176 (Title: Single particle cryo-EM dataset of the flexible and variable oligomeric state complex AP-1:Arf1:tetherin-HIV-NefData size: 115.4 Data #1: Dose weighted particle stack of AP1:Arf1:tetherin-HIV-Nef [picked particles - multiframe - unprocessed])  EMPIAR-10177 (Title: Single particle cryo-EM dataset of the flexible and variable oligomeric state complex AP-1:Arf1:tetherin-HIV-Nef EMPIAR-10177 (Title: Single particle cryo-EM dataset of the flexible and variable oligomeric state complex AP-1:Arf1:tetherin-HIV-NefData size: 34.0 Data #1: Dose weighted particle stack of AP1:Arf1:tetherin-HIV-Nef closed trimer [picked particles - multiframe - unprocessed])  EMPIAR-10178 (Title: Single particle cryo-EM dataset of the flexible and variable oligomeric state complex AP-1:Arf1:tetherin-HIV-Nef EMPIAR-10178 (Title: Single particle cryo-EM dataset of the flexible and variable oligomeric state complex AP-1:Arf1:tetherin-HIV-NefData size: 10.1 Data #1: Dose weight particle stack of AP1:Arf1:tetherin-HIV-Nef closed trimer after monomeric subunit extraction using LocalRec [picked particles - multiframe - processed]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_7455.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7455.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of the cargo bound AP-1:Arf1:tetherin-Nef monomer | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.067 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
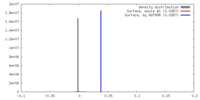
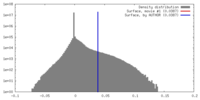
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Monomer of AP-1:Arf1:Tetherin-Nef
| Entire | Name: Monomer of AP-1:Arf1:Tetherin-Nef |
|---|---|
| Components |
|
-Supramolecule #1: Monomer of AP-1:Arf1:Tetherin-Nef
| Supramolecule | Name: Monomer of AP-1:Arf1:Tetherin-Nef / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 700 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 20 mM Tris at pH 8.0, 200 mM NaCl, 5 mM MgCl2, and 0.5 mM TCEP |
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 300 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 294 K / Instrument: FEI VITROBOT MARK IV |
| Details | AP-1:Arf1:Tetherin-Nef |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Dimensions - Width: 3712 pixel / Digitization - Dimensions - Height: 3840 pixel / Digitization - Frames/image: 3-38 / Number grids imaged: 1 / Number real images: 2200 / Average exposure time: 9.5 sec. / Average electron dose: 62.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 46849 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.6 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.75 µm / Nominal magnification: 22500 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)