[English] 日本語
 Yorodumi
Yorodumi- EMDB-7343: Ebola nucleoprotein nucleocapsid-like assembly and the asymmetric unit -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7343 | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Ebola nucleoprotein nucleocapsid-like assembly and the asymmetric unit | ||||||||||||||||||||||||
 Map data Map data | Ebola nucleoprotein nucleocapsid-like assembly | ||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||
 Keywords Keywords | Ebola / nucleoprotein / nucleocapsid / helical reconstruction / RNA BINDING PROTEIN | ||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationviral RNA genome packaging / helical viral capsid / viral nucleocapsid / host cell cytoplasm / ribonucleoprotein complex / RNA binding Similarity search - Function | ||||||||||||||||||||||||
| Biological species |  | ||||||||||||||||||||||||
| Method | helical reconstruction / cryo EM / Resolution: 5.8 Å | ||||||||||||||||||||||||
 Authors Authors | Su Z / Wu C / Pintilie GD / Chiu W / Amarasinghe GK / Leung DW | ||||||||||||||||||||||||
| Funding support |  United States, 7 items United States, 7 items
| ||||||||||||||||||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2015 Journal: Cell Rep / Year: 2015Title: An Intrinsically Disordered Peptide from Ebola Virus VP35 Controls Viral RNA Synthesis by Modulating Nucleoprotein-RNA Interactions. Authors: Daisy W Leung / Dominika Borek / Priya Luthra / Jennifer M Binning / Manu Anantpadma / Gai Liu / Ian B Harvey / Zhaoming Su / Ariel Endlich-Frazier / Juanli Pan / Reed S Shabman / Wah Chiu / ...Authors: Daisy W Leung / Dominika Borek / Priya Luthra / Jennifer M Binning / Manu Anantpadma / Gai Liu / Ian B Harvey / Zhaoming Su / Ariel Endlich-Frazier / Juanli Pan / Reed S Shabman / Wah Chiu / Robert A Davey / Zbyszek Otwinowski / Christopher F Basler / Gaya K Amarasinghe /  Abstract: During viral RNA synthesis, Ebola virus (EBOV) nucleoprotein (NP) alternates between an RNA-template-bound form and a template-free form to provide the viral polymerase access to the RNA template. In ...During viral RNA synthesis, Ebola virus (EBOV) nucleoprotein (NP) alternates between an RNA-template-bound form and a template-free form to provide the viral polymerase access to the RNA template. In addition, newly synthesized NP must be prevented from indiscriminately binding to noncognate RNAs. Here, we investigate the molecular bases for these critical processes. We identify an intrinsically disordered peptide derived from EBOV VP35 (NPBP, residues 20-48) that binds NP with high affinity and specificity, inhibits NP oligomerization, and releases RNA from NP-RNA complexes in vitro. The structure of the NPBP/ΔNPNTD complex, solved to 3.7 Å resolution, reveals how NPBP peptide occludes a large surface area that is important for NP-NP and NP-RNA interactions and for viral RNA synthesis. Together, our results identify a highly conserved viral interface that is important for EBOV replication and can be targeted for therapeutic development. | ||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7343.map.gz emd_7343.map.gz | 11.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7343-v30.xml emd-7343-v30.xml emd-7343.xml emd-7343.xml | 19.5 KB 19.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7343.png emd_7343.png | 204.8 KB | ||
| Filedesc metadata |  emd-7343.cif.gz emd-7343.cif.gz | 6.5 KB | ||
| Others |  emd_7343_additional.map.gz emd_7343_additional.map.gz emd_7343_additional_1.map.gz emd_7343_additional_1.map.gz | 117.5 KB 117.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7343 http://ftp.pdbj.org/pub/emdb/structures/EMD-7343 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7343 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7343 | HTTPS FTP |
-Related structure data
| Related structure data |  6c54MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_7343.map.gz / Format: CCP4 / Size: 70.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7343.map.gz / Format: CCP4 / Size: 70.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Ebola nucleoprotein nucleocapsid-like assembly | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.4 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Ebola nucleoprotein asymmetric unit
| File | emd_7343_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Ebola nucleoprotein asymmetric unit | ||||||||||||
| Projections & Slices |
| ||||||||||||
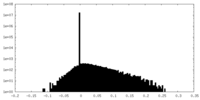
| Density Histograms |
-Additional map: Ebola nucleoprotein asymmetric unit
| File | emd_7343_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Ebola nucleoprotein asymmetric unit | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : eNP nucleocapsid-like assembly
| Entire | Name: eNP nucleocapsid-like assembly |
|---|---|
| Components |
|
-Supramolecule #1: eNP nucleocapsid-like assembly
| Supramolecule | Name: eNP nucleocapsid-like assembly / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Nucleoprotein
| Macromolecule | Name: Nucleoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 48.14259 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: LTAGLSVQQG IVRQRVIPVY QVNNLEEICQ LIIQAFEAGV DFQESADSFL LMLCLHHAYQ GDYKLFLESG AVKYLEGHGF RFEVKKRDG VKRLEELLPA VSSGKNIKRT LAAMPEEETT EANAGQFLSF ASLFLPKLVV GEKACLEKVQ RQIQVHAEQG L IQYPTAWQ ...String: LTAGLSVQQG IVRQRVIPVY QVNNLEEICQ LIIQAFEAGV DFQESADSFL LMLCLHHAYQ GDYKLFLESG AVKYLEGHGF RFEVKKRDG VKRLEELLPA VSSGKNIKRT LAAMPEEETT EANAGQFLSF ASLFLPKLVV GEKACLEKVQ RQIQVHAEQG L IQYPTAWQ SVGHMMVIFR LMRTNFLIKF LLIHQGMHMV AGHDANDAVI SNSVAQARFS GLLIVKTVLD HILQKTERGV RL HPLARTA KVKNEVNSFK AALSSLAKHG EYAPFARLLN LSGVNNLEHG LFPQLSAIAL GVATAHGSTL AGVNVGEQYQ QLR EAATEA EKQLQQYAES RELDHLGLDD QEKKILMNFH QKKNEISFQQ TNAMVTLRKE RLAKLTEAIT AASLPKTSGH YDDD DDIPF PGPINDDDNP GHQDDDPTDS QDTTIPDV UniProtKB: Nucleoprotein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Component - Concentration: 12.0 mM / Component - Formula: PBS / Component - Name: Phosphate-buffered saline / Details: 150 mM NaCl, 3 mM KCl, 10 mM Na2HPO4, 2 mM KH2PO4 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 40 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 3200FSC |
|---|---|
| Temperature | Min: 86.0 K / Max: 86.0 K |
| Specialist optics | Energy filter - Name: In-column Omega Filter / Energy filter - Lower energy threshold: 30 eV / Energy filter - Upper energy threshold: 30 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Digitization - Frames/image: 2-32 / Number grids imaged: 2 / Number real images: 1266 / Average exposure time: 8.0 sec. / Average electron dose: 24.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 4.7 mm / Nominal magnification: 30000 |
| Sample stage | Specimen holder model: JEOL 3200FSC CRYOHOLDER / Cooling holder cryogen: NITROGEN |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 2.65 Å Applied symmetry - Helical parameters - Δ&Phi: -8.53 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Resolution.type: BY AUTHOR / Resolution: 5.8 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 2-beta) / Number images used: 169526 |
|---|---|
| Segment selection | Number selected: 200685 |
| Startup model | Type of model: NONE |
| Final angle assignment | Type: NOT APPLICABLE / Software - Name: RELION (ver. 2-beta) |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 276 / Target criteria: Correlation coefficient |
|---|---|
| Output model |  PDB-6c54: |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)