+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7102 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | TPX2_micro decorated GMPCPP-microtubule | ||||||||||||||||||
 Map data Map data | TPX2_micro decorated GMPCPP-microtubule | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
| Biological species |   Homo sapiens (human) / Homo sapiens (human) /  | ||||||||||||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.6 Å | ||||||||||||||||||
 Authors Authors | Zhang R / Nogales E | ||||||||||||||||||
| Funding support |  United States, United States,  United Kingdom, 5 items United Kingdom, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Elife / Year: 2017 Journal: Elife / Year: 2017Title: Structural insight into TPX2-stimulated microtubule assembly. Authors: Rui Zhang / Johanna Roostalu / Thomas Surrey / Eva Nogales /   Abstract: During mitosis and meiosis, microtubule (MT) assembly is locally upregulated by the chromatin-dependent Ran-GTP pathway. One of its key targets is the MT-associated spindle assembly factor TPX2. The ...During mitosis and meiosis, microtubule (MT) assembly is locally upregulated by the chromatin-dependent Ran-GTP pathway. One of its key targets is the MT-associated spindle assembly factor TPX2. The molecular mechanism of how TPX2 stimulates MT assembly remains unknown because structural information about the interaction of TPX2 with MTs is lacking. Here, we determine the cryo-electron microscopy structure of a central region of TPX2 bound to the MT surface. TPX2 uses two flexibly linked elements ('ridge' and 'wedge') in a novel interaction mode to simultaneously bind across longitudinal and lateral tubulin interfaces. These MT-interacting elements overlap with the binding site of importins on TPX2. Fluorescence microscopy-based in vitro reconstitution assays reveal that this interaction mode is critical for MT binding and facilitates MT nucleation. Together, our results suggest a molecular mechanism of how the Ran-GTP gradient can regulate TPX2-dependent MT formation. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7102.map.gz emd_7102.map.gz | 156 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7102-v30.xml emd-7102-v30.xml emd-7102.xml emd-7102.xml | 15.5 KB 15.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7102.png emd_7102.png | 395.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7102 http://ftp.pdbj.org/pub/emdb/structures/EMD-7102 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7102 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7102 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_7102.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7102.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | TPX2_micro decorated GMPCPP-microtubule | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.33 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
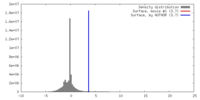
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : TPX2_micro decorated GMPCPP-microtubule
| Entire | Name: TPX2_micro decorated GMPCPP-microtubule |
|---|---|
| Components |
|
-Supramolecule #1: TPX2_micro decorated GMPCPP-microtubule
| Supramolecule | Name: TPX2_micro decorated GMPCPP-microtubule / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: tubulin
| Supramolecule | Name: tubulin / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: TPX2
| Supramolecule | Name: TPX2 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Spodoptera aff. frugiperda 2 RZ-2014 (butterflies/moths) Spodoptera aff. frugiperda 2 RZ-2014 (butterflies/moths)Recombinant strain: sf21 |
-Macromolecule #1: alpha tubulin
| Macromolecule | Name: alpha tubulin / type: other / ID: 1 / Classification: other |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MRECISIHVG QAGVQIGNAC WELYCLEHGI QPDGQMPSDK TIGGGDDSFN TFFSETGAGK HVPRAVFVDL EPTVIDEVRT GTYRQLFHPE QLITGKEDAA NNYARGHYTI GKEIIDLVLD RIRKLADQCT GLQGFLVFHS FGGGTGSGFT SLLMERLSVD YGKKSKLEFS ...String: MRECISIHVG QAGVQIGNAC WELYCLEHGI QPDGQMPSDK TIGGGDDSFN TFFSETGAGK HVPRAVFVDL EPTVIDEVRT GTYRQLFHPE QLITGKEDAA NNYARGHYTI GKEIIDLVLD RIRKLADQCT GLQGFLVFHS FGGGTGSGFT SLLMERLSVD YGKKSKLEFS IYPAPQVSTA VVEPYNSILT THTTLEHSDC AFMVDNEAIY DICRRNLDIE RPTYTNLNRL ISQIVSSITA SLRFDGALNV DLTEFQTNLV PYPRIHFPLA TYAPVISAEK AYHEQLSVAE ITNACFEPAN QMVKCDPRHG KYMACCLLYR GDVVPKDVNA AIATIKTKRS IQFVDWCPTG FKVGINYQPP TVVPGGDLAK VQRAVCMLSN TTAIAEAWAR LDHKFDLMYA KRAFVHWYVG EGMEEGEFSE AREDMAALEK DYEEVGVDSV EGEGEEEGEE Y |
-Macromolecule #2: beta tubulin
| Macromolecule | Name: beta tubulin / type: other / ID: 2 / Classification: other |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MREIVHIQAG QCGNQIGAKF WEVISDEHGI DPTGSYHGDS DLQLERINVY YNEAAGNKYV PRAILVDLEP GTMDSVRSGP FGQIFRPDNF VFGQSGAGNN WAKGHYTEGA ELVDSVLDVV RKESESCDCL QGFQLTHSLG GGTGSGMGTL LISKIREEYP DRIMNTFSVV ...String: MREIVHIQAG QCGNQIGAKF WEVISDEHGI DPTGSYHGDS DLQLERINVY YNEAAGNKYV PRAILVDLEP GTMDSVRSGP FGQIFRPDNF VFGQSGAGNN WAKGHYTEGA ELVDSVLDVV RKESESCDCL QGFQLTHSLG GGTGSGMGTL LISKIREEYP DRIMNTFSVV PSPKVSDTVV EPYNATLSVH QLVENTDETY CIDNEALYDI CFRTLKLTTP TYGDLNHLVS ATMSGVTTCL RFPGQLNADL RKLAVNMVPF PRLHFFMPGF APLTSRGSQQ YRALTVPELT QQMFDAKNMM AACDPRHGRY LTVAAVFRGR MSMKEVDEQM LNVQNKNSSY FVEWIPNNVK TAVCDIPPRG LKMSATFIGN STAIQELFKR ISEQFTAMFR RKAFLHWYTG EGMDEMEFTE AESNMNDLVS EYQQYQDATA DEQGEFEEEG EEDEA |
-Macromolecule #3: TPX2
| Macromolecule | Name: TPX2 / type: other / ID: 3 / Classification: other |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSQVKSSYSY DAPSDFINFS SLDDEGDTQN IDSWFEEKAN LENKLLGKNG TGGLFQGKTP LRKANLQQA IVTPLKPVDN TYYKEAEKEN LVEQSIPSNA CSSLEVEAAI SRKTPAQPQR R SLRLSAQK DLEQKEKHHV KMKAKRCATP VIIDEILPSK KMKVSNNKKK ...String: MSQVKSSYSY DAPSDFINFS SLDDEGDTQN IDSWFEEKAN LENKLLGKNG TGGLFQGKTP LRKANLQQA IVTPLKPVDN TYYKEAEKEN LVEQSIPSNA CSSLEVEAAI SRKTPAQPQR R SLRLSAQK DLEQKEKHHV KMKAKRCATP VIIDEILPSK KMKVSNNKKK PEEEGSAHQD TA EKNASSP EKAKGRHTVP CMPPAKQKFL KSTEEQELEK SMKMQQEVVE MRKKNEEFKK LAL AGIGQP VKKSVSQVTK SVDFHFRTDE RIKQHPKNQE EYKEVNFTSE LRKHPSSPAR VTKG CTIVK PFNLSQGKKR TFDETVSTYV PLAQQVEDFH KRTPNRYHLR SKKDDINLLP SKSSV TKIC RDPQTPVLQT KHRARAVTCK STAELEAEEL EKLQQYKFKA RELDPRILEG GPILPK KPP VKPPTEPIGF DLEIEKRIQE RESKKKTEDE HFEFHSRPCP TKILEDVVGV PEKKVLP IT VPKSPAFALK NRIRMPTKED EEEDEPVVIK AQPVPHYGVP FKPQIPEART VEICPFSF D SRDKERQLQK EKKIKELQKG EVPKFKALPL PHFDTINLPE KKVKNVTQIE PFCLETDRR GALKAQTWKH QLEEELRQQK EAACFKARPN TVISQEPFVP KKEKKSVAEG LSGSLVQEPF QLATEKRAK ERQELEKRMA EVEAQKAQQL EEARLQEEEQ KKEELARLRR ELVHKANPIR K YQGLEIKS SDQPLTVPVS PKFSTRFHC |
| Recombinant expression | Organism:  Spodoptera aff. frugiperda 2 RZ-2014 (butterflies/moths) Spodoptera aff. frugiperda 2 RZ-2014 (butterflies/moths) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Buffer | pH: 6.8 / Details: BRB80 |
|---|---|
| Grid | Model: C-flat-1.2/1.3 4C / Material: COPPER / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 288 K / Instrument: FEI VITROBOT MARK II / Details: blot for 4 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 4 / Number real images: 2000 / Average exposure time: 6.0 sec. / Average electron dose: 27.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: GATAN 626 SINGLE TILT LIQUID NITROGEN CRYO TRANSFER HOLDER Cooling holder cryogen: NITROGEN |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 9.04 Å Applied symmetry - Helical parameters - Δ&Phi: -25.74 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Algorithm: FOURIER SPACE / Resolution.type: BY AUTHOR / Resolution: 3.6 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: FREALIGN / Number images used: 85000 |
|---|---|
| CTF correction | Software - Name: CTFFIND |
| Final angle assignment | Type: NOT APPLICABLE / Software - Name: FREALIGN |
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)