+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Bacillus ytrEF vanadate trapped conformation | |||||||||
 Map data Map data | A main cryoEM map with sharpening factor of -101.5. The resolution of this map is 2.4 angstrom at a FSC threshold of 0.143. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ABC Transporter / vanadate / ATP / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmembrane protein complex / transmembrane transporter activity / transmembrane transport / ATP hydrolysis activity / ATP binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.5 Å | |||||||||
 Authors Authors | Yu P / Krah BS / Orlando MA / Orlando BJ | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2026 Journal: Structure / Year: 2026Title: Structural analysis of a Gram-positive type VII ABC transporter induced by cell wall-targeting antibiotics. Authors: Peixuan Yu / Bradon S Krah / Melanie A Orlando / Sundharraman Subramanian / Benjamin J Orlando /  Abstract: Bacteria utilize a variety of mechanisms to remodel the cell wall in response to environmental and antimicrobial stress. In the model organism Bacillus subtilis, the ytr operon encoding putative ATP- ...Bacteria utilize a variety of mechanisms to remodel the cell wall in response to environmental and antimicrobial stress. In the model organism Bacillus subtilis, the ytr operon encoding putative ATP-binding cassette (ABC) transporter(s) is highly upregulated in response to cell wall-targeting antibiotics. Here we show that the ytr operon encodes two distinct ABC transporters: YtrBCD and YtrEF. Using cryo-electron microscopy(cryo-EM), we determined the structures of YtrEF in nucleotide-free and ADP-vanadate bound states. The structures demonstrate that YtrEF adopts a type VII ABC transporter fold. Nucleotide binding induced conformational changes that propagate from the cytosolic region through the transmembrane helices to ultimately reorient the extracellular domains. Extended bacterial growth assays and suppressor mutation identification indicated that YtrEF contributes to alteration of colony morphology. These findings establish YtrEF as a type VII ABC transporter that is induced by cell wall-targeting antibiotics and a new avenue to phenotypically assess the ytr operon. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_70266.map.gz emd_70266.map.gz | 118 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-70266-v30.xml emd-70266-v30.xml emd-70266.xml emd-70266.xml | 30.7 KB 30.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_70266_fsc.xml emd_70266_fsc.xml | 10.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_70266.png emd_70266.png | 57.8 KB | ||
| Filedesc metadata |  emd-70266.cif.gz emd-70266.cif.gz | 8 KB | ||
| Others |  emd_70266_additional_1.map.gz emd_70266_additional_1.map.gz emd_70266_additional_2.map.gz emd_70266_additional_2.map.gz emd_70266_half_map_1.map.gz emd_70266_half_map_1.map.gz emd_70266_half_map_2.map.gz emd_70266_half_map_2.map.gz | 109.7 MB 62.6 MB 115.9 MB 115.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-70266 http://ftp.pdbj.org/pub/emdb/structures/EMD-70266 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-70266 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-70266 | HTTPS FTP |
-Related structure data
| Related structure data |  9o9yMC  9o9xC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_70266.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_70266.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | A main cryoEM map with sharpening factor of -101.5. The resolution of this map is 2.4 angstrom at a FSC threshold of 0.143. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.886 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: A cryoEM map derived from DeepEMhancer algorithm.
| File | emd_70266_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | A cryoEM map derived from DeepEMhancer algorithm. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: unsharpened cryoEM map
| File | emd_70266_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened cryoEM map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: unsharpened cryoEM map A
| File | emd_70266_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened cryoEM map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: unsharpened cryoEM map B
| File | emd_70266_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened cryoEM map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Bacillus ABC transporter ytrEF in vanadate-bound conformation
| Entire | Name: Bacillus ABC transporter ytrEF in vanadate-bound conformation |
|---|---|
| Components |
|
-Supramolecule #1: Bacillus ABC transporter ytrEF in vanadate-bound conformation
| Supramolecule | Name: Bacillus ABC transporter ytrEF in vanadate-bound conformation type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 150 KDa |
-Macromolecule #1: ABC transporter ATP-binding protein YtrE
| Macromolecule | Name: ABC transporter ATP-binding protein YtrE / type: protein_or_peptide / ID: 1 Details: The construct has a HIS tag and a thrombin cleavage site in the N terminal of ytrE. Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: Strain 168 |
| Molecular weight | Theoretical: 27.664412 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSSHHHHHH SSGLVPRGSH MIDVQHIDHS FTIGKKGREN EVPVLKDVSL SVAKGEIACI VGRSGSGKST LLNLISGYIS PTKGRIVIN GTDVTGFNEK EWAQFRLDHF GFIFQSFQLI PGLTTYENVE MPLALKGIKP SERKQKVQDM LKRVGLENHA A HYPNELSG ...String: MGSSHHHHHH SSGLVPRGSH MIDVQHIDHS FTIGKKGREN EVPVLKDVSL SVAKGEIACI VGRSGSGKST LLNLISGYIS PTKGRIVIN GTDVTGFNEK EWAQFRLDHF GFIFQSFQLI PGLTTYENVE MPLALKGIKP SERKQKVQDM LKRVGLENHA A HYPNELSG GQQQRVSIAR ALILNPSIIL ADEPTGSLDS ETEHEVLELI QQLNRERGIT FVIITHDDEV ASIGHSKFQL HD GVLKGGI TVEV UniProtKB: ABC transporter ATP-binding protein YtrE |
-Macromolecule #2: ABC transporter permease YtrF
| Macromolecule | Name: ABC transporter permease YtrF / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 48.477973 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MRFKDQVHFI RRNMKKNRLR VFMTILATTM ACAFLVVLSS VGFGIQKTIT DMTMSQQIVT KVSVMGKEGD KPIKKADLEK YDHVRSVVE RTQVYEPNKA TLGNRTNESS NLIFTNMNDE LKANMELEKG RVAKSENEIV VGYDFAKRLL TKKESEEYNK K IEEAKGNP ...String: MRFKDQVHFI RRNMKKNRLR VFMTILATTM ACAFLVVLSS VGFGIQKTIT DMTMSQQIVT KVSVMGKEGD KPIKKADLEK YDHVRSVVE RTQVYEPNKA TLGNRTNESS NLIFTNMNDE LKANMELEKG RVAKSENEIV VGYDFAKRLL TKKESEEYNK K IEEAKGNP EDIKEPKGYT KDILNKTIEL SVSKTDSKTG DVTKTKTYDF KIVGITKKPS QDWMEDSNIF ISDQFKKDFS EF LDFKGGN VETNIGVFAD KFENVEQLTN DLTDDGYYVT SVTTELEGAN TFFMVFKIGL IFVGCIAVII SAIGIFNTMT MAV TERTQE IGIMKAIGAS PSIIRRMFLM ESAYIGILGC VIGIIISYGV SYLVNLAVPM ILAATSGGDA GDLNYTFSYI PASL VIIAV VICGGVAVIS GMNPARKATK TNVLTALRRE L UniProtKB: ABC transporter permease YtrF |
-Macromolecule #3: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 2 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #4: VANADATE ION
| Macromolecule | Name: VANADATE ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: VO4 |
|---|---|
| Molecular weight | Theoretical: 114.939 Da |
| Chemical component information | 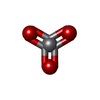 ChemComp-VN3: |
-Macromolecule #5: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #6: UNKNOWN LIGAND
| Macromolecule | Name: UNKNOWN LIGAND / type: ligand / ID: 6 / Number of copies: 2 / Formula: UNL |
|---|---|
| Chemical component information | 
ChemComp-UNL: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||||||||
| Grid | Model: Quantifoil / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. / Pretreatment - Atmosphere: AIR | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV | |||||||||||||||
| Details | The ytrEF in nucleotide-free conformation is mono disperse and pure before plunge-frozen onto the cryoEM grids. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris |
| Image recording | Film or detector model: TFS FALCON 4i (4k x 4k) / Number grids imaged: 1 / Number real images: 11530 / Average electron dose: 44.34 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Details | Refinement in phenix.real_space_refine |
| Refinement | Space: REAL / Protocol: OTHER |
| Output model |  PDB-9o9y: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















































