+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6951 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Hepatitis B virus spherical subviral particle | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |   Hepatitis B virus Hepatitis B virus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 30.1 Å | |||||||||
 Authors Authors | Cao J / Luo S / Zhang J / Zhu P | |||||||||
 Citation Citation |  Journal: Virus Res / Year: 2019 Journal: Virus Res / Year: 2019Title: Cryo-EM structure of native spherical subviral particles isolated from HBV carriers. Authors: Jianhao Cao / Junchang Zhang / Yanmeng Lu / Shuhong Luo / Jingqiang Zhang / Ping Zhu /  Abstract: Hepatitis B virus (HBV) contains 3 types of particles, i.e., 22-nm-diameter spherical and tubular subviral particles (SVPs) and 44-nm-diameter Dane particles. The SVPs are non-infectious and present ...Hepatitis B virus (HBV) contains 3 types of particles, i.e., 22-nm-diameter spherical and tubular subviral particles (SVPs) and 44-nm-diameter Dane particles. The SVPs are non-infectious and present strong immunogenicity, while Dane particles are infectious. In this study, we isolated spherical SVPs from HBV carriers' sera and determined their 3D structure at the resolution of ∼30 Å by cryo-electron microscopy (cryo-EM) single-particle reconstruction. Our cryo-EM structure suggests that the native HBV spherical SVP is irregularly organized, where spike-like features are arranged in a crystalline-like pattern on the surface. Strikingly, the hepatitis B surface antigen (HBsAg) in the native spherical SVPs folds as protrusions on the surface, as those on the native tubular SVPs and Dane particles, but is largely different from that in the recombinant octahedral SVPs. These results suggest a universal folding shape of HBsAg on the native HBV viral and subviral particles. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6951.map.gz emd_6951.map.gz | 20.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6951-v30.xml emd-6951-v30.xml emd-6951.xml emd-6951.xml | 7.2 KB 7.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_6951.png emd_6951.png | 75.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6951 http://ftp.pdbj.org/pub/emdb/structures/EMD-6951 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6951 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6951 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_6951.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6951.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.5 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
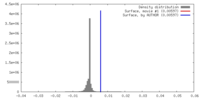
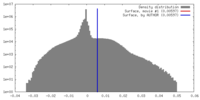
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Hepatitis B virus
| Entire | Name:   Hepatitis B virus Hepatitis B virus |
|---|---|
| Components |
|
-Supramolecule #1: Hepatitis B virus
| Supramolecule | Name: Hepatitis B virus / type: virus / ID: 1 / Parent: 0 / NCBI-ID: 10407 / Sci species name: Hepatitis B virus / Virus type: VIRION / Virus isolate: SEROCOMPLEX / Virus enveloped: Yes / Virus empty: Yes |
|---|
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Sugar embedding | Material: ice |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: DIRECT ELECTRON DE-20 (5k x 3k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 30.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 5478 |
|---|---|
| Initial angle assignment | Type: NOT APPLICABLE |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)