+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6923 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | M. smegmatis P/P state 30S ribosomal subunit | |||||||||
 Map data Map data | 30S masked map of translating-state ribosome of M. smegmatis | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | translating-state / ribosome / complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationribosomal small subunit assembly / ribosome biogenesis / ribosomal small subunit biogenesis / small ribosomal subunit / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / tRNA binding / rRNA binding / structural constituent of ribosome / ribosome ...ribosomal small subunit assembly / ribosome biogenesis / ribosomal small subunit biogenesis / small ribosomal subunit / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / tRNA binding / rRNA binding / structural constituent of ribosome / ribosome / translation / ribonucleoprotein complex / mRNA binding / RNA binding / zinc ion binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Mycobacterium smegmatis str. MC2 155 (bacteria) / Mycobacterium smegmatis str. MC2 155 (bacteria) /   Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria) Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Mishra S / Ahmed T | |||||||||
| Funding support |  Singapore, 1 items Singapore, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Rep / Year: 2018 Journal: Sci Rep / Year: 2018Title: Structures of Mycobacterium smegmatis 70S ribosomes in complex with HPF, tmRNA, and P-tRNA. Authors: Satabdi Mishra / Tofayel Ahmed / Anu Tyagi / Jian Shi / Shashi Bhushan /  Abstract: Ribosomes are the dynamic protein synthesis machineries of the cell. They may exist in different functional states in the cell. Therefore, it is essential to have structural information on these ...Ribosomes are the dynamic protein synthesis machineries of the cell. They may exist in different functional states in the cell. Therefore, it is essential to have structural information on these different functional states of ribosomes to understand their mechanism of action. Here, we present single particle cryo-EM reconstructions of the Mycobacterium smegmatis 70S ribosomes in the hibernating state (with HPF), trans-translating state (with tmRNA), and the P/P state (with P-tRNA) resolved to 4.1, 12.5, and 3.4 Å, respectively. A comparison of the P/P state with the hibernating state provides possible functional insights about the Mycobacteria-specific helix H54a rRNA segment. Interestingly, densities for all the four OB domains of bS1 protein is visible in the hibernating 70S ribosome displaying the molecular details of bS1-70S interactions. Our structural data shows a Mycobacteria-specific H54a-bS1 interaction which seems to prevent subunit dissociation and degradation during hibernation without the formation of 100S dimer. This indicates a new role of bS1 protein in 70S protection during hibernation in Mycobacteria in addition to its conserved function during translation initiation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6923.map.gz emd_6923.map.gz | 25.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6923-v30.xml emd-6923-v30.xml emd-6923.xml emd-6923.xml | 39.5 KB 39.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_6923.png emd_6923.png | 106.6 KB | ||
| Filedesc metadata |  emd-6923.cif.gz emd-6923.cif.gz | 9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6923 http://ftp.pdbj.org/pub/emdb/structures/EMD-6923 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6923 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6923 | HTTPS FTP |
-Related structure data
| Related structure data |  5zeuMC  6920C  6921C  6922C  6925C  5zebC  5zepC  5zetC  5zeyC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6923.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6923.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 30S masked map of translating-state ribosome of M. smegmatis | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
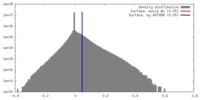
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Translating-state 30S ribosome structure from M.smegmatis
+Supramolecule #1: Translating-state 30S ribosome structure from M.smegmatis
+Supramolecule #2: 30S ribosome
+Supramolecule #3: P-tRNAfMet
+Macromolecule #1: 16S rRNA
+Macromolecule #15: P-tRNAfMet
+Macromolecule #2: 30S ribosomal protein S3
+Macromolecule #3: 30S ribosomal protein S5
+Macromolecule #4: 30S ribosomal protein S7
+Macromolecule #5: 30S ribosomal protein S8
+Macromolecule #6: 30S ribosomal protein S9
+Macromolecule #7: 30S ribosomal protein S10
+Macromolecule #8: 30S ribosomal protein S11
+Macromolecule #9: 30S ribosomal protein S12
+Macromolecule #10: 30S ribosomal protein S15
+Macromolecule #11: 30S ribosomal protein S17
+Macromolecule #12: 30S ribosomal protein S18 2
+Macromolecule #13: 30S ribosomal protein S19
+Macromolecule #14: 30S ribosomal protein S20
+Macromolecule #16: 30S ribosomal protein S14 type Z
+Macromolecule #17: 30S ribosomal protein S2
+Macromolecule #18: 30S ribosomal protein S4
+Macromolecule #19: 30S ribosomal protein S6
+Macromolecule #20: 30S ribosomal protein S13
+Macromolecule #21: 30S ribosomal protein S16
+Macromolecule #22: Conserved domain protein
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Material: COPPER |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Average electron dose: 1.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)