+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-6876 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Cryo-EM structure of Methanoccus maripaludis archaellum | |||||||||
 マップデータ マップデータ | Methanococcus maripaludis archaellum cryo-EM reconstruction | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | Archaellum / archea / flagellum / metal binding / motility / helical / PROTEIN FIBRIL | |||||||||
| 機能・相同性 | archaeal-type flagellum / Flagellin, archaea / Archaeal-type flagellin / Flagellin/pilin, N-terminal / archaeal or bacterial-type flagellum-dependent cell motility / structural molecule activity / Flagellin 機能・相同性情報 機能・相同性情報 | |||||||||
| 生物種 |  Methanococcus maripaludis S2 (古細菌) / Methanococcus maripaludis S2 (古細菌) /  Methanococcus maripaludis (strain S2 / LL) (古細菌) Methanococcus maripaludis (strain S2 / LL) (古細菌) | |||||||||
| 手法 | らせん対称体再構成法 / クライオ電子顕微鏡法 / 解像度: 4.0 Å | |||||||||
 データ登録者 データ登録者 | Meshcheryakov VA / Shibata S / Schreiber MT / Villar-Briones A / Jarrell KF / Aizawa S / Wolf M / Kurumizaka H | |||||||||
| 資金援助 |  日本, 2件 日本, 2件
| |||||||||
 引用 引用 |  ジャーナル: EMBO Rep / 年: 2019 ジャーナル: EMBO Rep / 年: 2019タイトル: High-resolution archaellum structure reveals a conserved metal-binding site. 著者: Vladimir A Meshcheryakov / Satoshi Shibata / Makoto Tokoro Schreiber / Alejandro Villar-Briones / Kenneth F Jarrell / Shin-Ichi Aizawa / Matthias Wolf /   要旨: Many archaea swim by means of archaella. While the archaellum is similar in function to its bacterial counterpart, its structure, composition, and evolution are fundamentally different. Archaella are ...Many archaea swim by means of archaella. While the archaellum is similar in function to its bacterial counterpart, its structure, composition, and evolution are fundamentally different. Archaella are related to archaeal and bacterial type IV pili. Despite recent advances, our understanding of molecular processes governing archaellum assembly and stability is still incomplete. Here, we determine the structures of archaella by X-ray crystallography and cryo-EM The crystal structure of FlaB1 is the first and only crystal structure of any archaellin to date at a resolution of 1.5 Å, which is put into biological context by a cryo-EM reconstruction from archaella at 4 Å resolution created with helical single-particle analysis. Our results indicate that the archaellum is predominantly composed of FlaB1. We identify N-linked glycosylation by cryo-EM and mass spectrometry. The crystal structure reveals a highly conserved metal-binding site, which is validated by mass spectrometry and electron energy-loss spectroscopy. We show that the metal-binding site, which appears to be a widespread property of archaellin, is required for filament integrity. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_6876.map.gz emd_6876.map.gz | 2.7 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-6876-v30.xml emd-6876-v30.xml emd-6876.xml emd-6876.xml | 14.3 KB 14.3 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_6876.png emd_6876.png | 119.9 KB | ||
| Filedesc metadata |  emd-6876.cif.gz emd-6876.cif.gz | 5.8 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6876 http://ftp.pdbj.org/pub/emdb/structures/EMD-6876 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6876 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6876 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_6876.map.gz / 形式: CCP4 / 大きさ: 8 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_6876.map.gz / 形式: CCP4 / 大きさ: 8 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Methanococcus maripaludis archaellum cryo-EM reconstruction | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.41 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
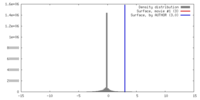
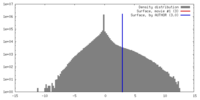
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
-全体 : M.maripaludis archaellin FlaB1 filament
| 全体 | 名称: M.maripaludis archaellin FlaB1 filament |
|---|---|
| 要素 |
|
-超分子 #1: M.maripaludis archaellin FlaB1 filament
| 超分子 | 名称: M.maripaludis archaellin FlaB1 filament / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: all |
|---|---|
| 由来(天然) | 生物種:  Methanococcus maripaludis S2 (古細菌) Methanococcus maripaludis S2 (古細菌) |
-分子 #1: Flagellin
| 分子 | 名称: Flagellin / タイプ: protein_or_peptide / ID: 1 / コピー数: 18 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Methanococcus maripaludis (strain S2 / LL) (古細菌) Methanococcus maripaludis (strain S2 / LL) (古細菌)株: S2 / LL |
| 分子量 | 理論値: 21.639359 KDa |
| 配列 | 文字列: MKITEFMKSK KGASGIGTLI VFIAMVLVAA VAASVLINTS GFLQQKASTT GKESTEQVAS GLLINGITGS VGTSDVKLLA IYLAPNAGS SAIDLAQTKV MLDYNGKSVV LGYGGNQDMS SGNSSVFSND TGATATTFQV NILQDYDDSA VDNAVINKGD A VALIVDVN ...文字列: MKITEFMKSK KGASGIGTLI VFIAMVLVAA VAASVLINTS GFLQQKASTT GKESTEQVAS GLLINGITGS VGTSDVKLLA IYLAPNAGS SAIDLAQTKV MLDYNGKSVV LGYGGNQDMS SGNSSVFSND TGATATTFQV NILQDYDDSA VDNAVINKGD A VALIVDVN ASFAGEIPER TAISGKVQPE FGAPGVISFT TPASYTTTLV ELQ UniProtKB: Flagellin |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | らせん対称体再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | .8 mg/mL |
|---|---|
| 緩衝液 | pH: 7 / 構成要素 - 式: H2O / 構成要素 - 名称: Water |
| グリッド | モデル: Quantifoil R1.2/1.3 / 材質: COPPER / メッシュ: 200 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: HOLEY / 支持フィルム - Film thickness: 100 / 前処理 - タイプ: PLASMA CLEANING / 前処理 - 時間: 15 sec. / 前処理 - 雰囲気: OTHER / 前処理 - 気圧: 9.33 kPa / 詳細: Gatan Solarus |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 289 K / 装置: FEI VITROBOT MARK IV / 詳細: 3 second blot, 3.5uL. |
| 詳細 | sample was monodisperse |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 温度 | 最低: 77.0 K / 最高: 100.0 K |
| アライメント法 | Coma free - Residual tilt: 0.1 mrad |
| 特殊光学系 | エネルギーフィルター - 名称: GIF Quantum LS エネルギーフィルター - エネルギー下限: 0 eV エネルギーフィルター - エネルギー上限: 20 eV |
| 詳細 | nanoprobe, parallel beam illumination |
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 検出モード: SUPER-RESOLUTION / デジタル化 - サイズ - 横: 7676 pixel / デジタル化 - サイズ - 縦: 7420 pixel / デジタル化 - 画像ごとのフレーム数: 1-48 / 撮影したグリッド数: 1 / 実像数: 2000 / 平均露光時間: 12.0 sec. / 平均電子線量: 96.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | C2レンズ絞り径: 50.0 µm / 最大 デフォーカス(補正後): -2.5 µm / 最小 デフォーカス(補正後): -1.5 µm / 倍率(補正後): 47619 / 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 2.7 mm / 最大 デフォーカス(公称値): -2.5 µm / 最小 デフォーカス(公称値): -1.5 µm / 倍率(公称値): 105000 |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER ホルダー冷却材: NITROGEN |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
+ 画像解析
画像解析
-原子モデル構築 1
| 精密化 | 空間: REAL / プロトコル: FLEXIBLE FIT |
|---|---|
| 得られたモデル |  PDB-5z1l: |
 ムービー
ムービー コントローラー
コントローラー











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)