+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6791 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of p300-p53 protein complex | |||||||||
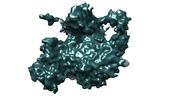
 Map data Map data | Cryo-EM map of p300-p53 complex. p53 protein is a homotetrameric tumor suppresor. Transcriptional coactivator p300 is a multidomain protein with HAT (histone acetyltransferase) activity. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | transcription factor / autoacetylation / allosteric interaction / catalytically active form / TRANSCRIPTION | |||||||||
| Function / homology |  Function and homology information Function and homology informationbehavioral defense response / negative regulation of protein oligomerization / peptidyl-lysine propionylation / histone lactyltransferase (CoA-dependent) activity / peptidyl-lysine crotonylation / peptidyl-lysine butyrylation / histone butyryltransferase activity / swimming / histone H3K122 acetyltransferase activity / peptide butyryltransferase activity ...behavioral defense response / negative regulation of protein oligomerization / peptidyl-lysine propionylation / histone lactyltransferase (CoA-dependent) activity / peptidyl-lysine crotonylation / peptidyl-lysine butyrylation / histone butyryltransferase activity / swimming / histone H3K122 acetyltransferase activity / peptide butyryltransferase activity / regulation of tubulin deacetylation / histone H2B acetyltransferase activity / internal protein amino acid acetylation / peptide 2-hydroxyisobutyryltransferase activity / histone crotonyltransferase activity / protein propionyltransferase activity / NOTCH2 intracellular domain regulates transcription / thigmotaxis / L-lysine N-acetyltransferase activity, acting on acetyl phosphate as donor / positive regulation of TORC2 signaling / internal peptidyl-lysine acetylation / histone H4 acetyltransferase activity / cellular response to L-leucine / histone H3 acetyltransferase activity / NFE2L2 regulating ER-stress associated genes / acetylation-dependent protein binding / Activation of the TFAP2 (AP-2) family of transcription factors / NFE2L2 regulating inflammation associated genes / NGF-stimulated transcription / histone H3K18 acetyltransferase activity / N-terminal peptidyl-lysine acetylation / histone H3K27 acetyltransferase activity / LRR FLII-interacting protein 1 (LRRFIP1) activates type I IFN production / NFE2L2 regulates pentose phosphate pathway genes / STAT3 nuclear events downstream of ALK signaling / Polo-like kinase mediated events / NFE2L2 regulating MDR associated enzymes / host-mediated activation of viral transcription / negative regulation of helicase activity / Loss of function of TP53 in cancer due to loss of tetramerization ability / Regulation of TP53 Expression / TGFBR3 expression / signal transduction by p53 class mediator / negative regulation of G1 to G0 transition / negative regulation of glucose catabolic process to lactate via pyruvate / regulation of androgen receptor signaling pathway / Transcriptional activation of cell cycle inhibitor p21 / regulation of intrinsic apoptotic signaling pathway by p53 class mediator / negative regulation of pentose-phosphate shunt / ATP-dependent DNA/DNA annealing activity / Activation of NOXA and translocation to mitochondria / regulation of mitochondrion organization / regulation of cell cycle G2/M phase transition / regulation of fibroblast apoptotic process / intrinsic apoptotic signaling pathway in response to hypoxia / oligodendrocyte apoptotic process / negative regulation of miRNA processing / positive regulation of thymocyte apoptotic process / oxidative stress-induced premature senescence / regulation of tissue remodeling / positive regulation of mitochondrial membrane permeability / mRNA transcription / positive regulation of programmed necrotic cell death / Regulation of gene expression in late stage (branching morphogenesis) pancreatic bud precursor cells / Regulation of FOXO transcriptional activity by acetylation / bone marrow development / RUNX3 regulates NOTCH signaling / circadian behavior / T cell proliferation involved in immune response / regulation of mitochondrial membrane permeability involved in apoptotic process / NOTCH4 Intracellular Domain Regulates Transcription / germ cell nucleus / Nuclear events mediated by NFE2L2 / Regulation of NFE2L2 gene expression / RUNX3 regulates CDKN1A transcription / Regulation of gene expression by Hypoxia-inducible Factor / face morphogenesis / glucose catabolic process to lactate via pyruvate / TP53 Regulates Transcription of Death Receptors and Ligands / TP53 regulates transcription of additional cell cycle genes whose exact role in the p53 pathway remain uncertain / Activation of PUMA and translocation to mitochondria / platelet formation / regulation of glycolytic process / NOTCH3 Intracellular Domain Regulates Transcription / regulation of DNA damage response, signal transduction by p53 class mediator / TRAF6 mediated IRF7 activation / histone deacetylase regulator activity / NFE2L2 regulating tumorigenic genes / NFE2L2 regulating anti-oxidant/detoxification enzymes / megakaryocyte development / negative regulation of glial cell proliferation / Regulation of TP53 Activity through Association with Co-factors / negative regulation of neuroblast proliferation / protein-lysine-acetyltransferase activity / acyltransferase activity / nuclear androgen receptor binding / STAT family protein binding / mitochondrial DNA repair / T cell lineage commitment / Formation of Senescence-Associated Heterochromatin Foci (SAHF) Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
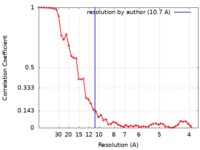
| Method | single particle reconstruction / cryo EM / Resolution: 10.7 Å | |||||||||
 Authors Authors | Ghosh R / Roy S / Sengupta J | |||||||||
 Citation Citation |  Journal: Biochemistry / Year: 2019 Journal: Biochemistry / Year: 2019Title: Tumor Suppressor p53-Mediated Structural Reorganization of the Transcriptional Coactivator p300. Authors: Raka Ghosh / Stephanie Kaypee / Manidip Shasmal / Tapas K Kundu / Siddhartha Roy / Jayati Sengupta /  Abstract: Transcriptional coactivator p300, a critical player in eukaryotic gene regulation, primarily functions as a histone acetyltransferase (HAT). It is also an important player in acetylation of a number ...Transcriptional coactivator p300, a critical player in eukaryotic gene regulation, primarily functions as a histone acetyltransferase (HAT). It is also an important player in acetylation of a number of nonhistone proteins, p53 being the most prominent one. Recruitment of p300 to p53 is pivotal in the regulation of p53-dependent genes. Emerging evidence suggests that p300 adopts an active conformation upon binding to the tetrameric p53, resulting in its enhanced acetylation activity. As a modular protein, p300 consists of multiple well-defined domains, where the structured domains are interlinked with unstructured linker regions. A crystal structure of the central domain of p300 encompassing Bromo, RING, PHD, and HAT domains demonstrates a compact module, where the HAT active site stays occluded by the RING domain. However, although p300 has a significant role in mediating the transcriptional activity of p53, only a few structural details on the complex of these two full-length proteins are available. Here, we present a cryo-electron microscopy (cryo-EM) study on the p300-p53 complex. The three-dimensional cryo-EM density map of the p300-p53 complex, when compared to the cryo-EM map of free p300, revealed that substantial change in the relative arrangement of Bromo and HAT domains occurs upon complex formation, which is likely required for exposing HAT active site and subsequent acetyltransferase activity. Our observation correlates well with previous studies showing that the presence of Bromodomain is obligatory for effective acetyltransferase activity of HAT. Thus, our result sheds new light on the mechanism whereby p300, following binding with p53, gets activated. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6791.map.gz emd_6791.map.gz | 7.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6791-v30.xml emd-6791-v30.xml emd-6791.xml emd-6791.xml | 15.1 KB 15.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_6791_fsc.xml emd_6791_fsc.xml | 5.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_6791.png emd_6791.png | 153.5 KB | ||
| Filedesc metadata |  emd-6791.cif.gz emd-6791.cif.gz | 6.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6791 http://ftp.pdbj.org/pub/emdb/structures/EMD-6791 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6791 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6791 | HTTPS FTP |
-Validation report
| Summary document |  emd_6791_validation.pdf.gz emd_6791_validation.pdf.gz | 507.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_6791_full_validation.pdf.gz emd_6791_full_validation.pdf.gz | 507.4 KB | Display | |
| Data in XML |  emd_6791_validation.xml.gz emd_6791_validation.xml.gz | 8.4 KB | Display | |
| Data in CIF |  emd_6791_validation.cif.gz emd_6791_validation.cif.gz | 10.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6791 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6791 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6791 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6791 | HTTPS FTP |
-Related structure data
| Related structure data |  5xzcMC  6792C  6k4nC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6791.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6791.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM map of p300-p53 complex. p53 protein is a homotetrameric tumor suppresor. Transcriptional coactivator p300 is a multidomain protein with HAT (histone acetyltransferase) activity. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.89 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
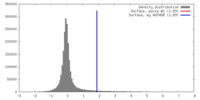
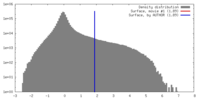
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : p300-p53 complex
| Entire | Name: p300-p53 complex |
|---|---|
| Components |
|
-Supramolecule #1: p300-p53 complex
| Supramolecule | Name: p300-p53 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Proteins were purified separately and then complex was made for cryo-EM. |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Histone acetyltransferase p300
| Macromolecule | Name: Histone acetyltransferase p300 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: histone acetyltransferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 72.100062 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: KKIFKPEELR QALMPTLEAL YRQDPESLPF RQPVDPQLLG IPDYFDIVKS PMDLSTIKRK LDTGQYQEPW QYVDDIWLMF NNAWLYNRK TSRVYKYCSK LSEVFEQEID PVMQSLGYCC GRKLEFSPQT LCCYGKQLCT IPRDATYYSY QNRYHFCEKC F NEIQGESV ...String: KKIFKPEELR QALMPTLEAL YRQDPESLPF RQPVDPQLLG IPDYFDIVKS PMDLSTIKRK LDTGQYQEPW QYVDDIWLMF NNAWLYNRK TSRVYKYCSK LSEVFEQEID PVMQSLGYCC GRKLEFSPQT LCCYGKQLCT IPRDATYYSY QNRYHFCEKC F NEIQGESV SLGDDPSQPQ TTINKEQFSK RKNDTLDPEL FVECTECGRK MHQICVLHHE IIWPAGFVCD GCLKKSARTR KE NKFSAKR LPSTRLGTFL ENRVNDFLRR QNHPESGEVT VRVVHASDKT VEVKPGMKAR FVDSGEMAES FPYRTKALFA FEE IDGVDL CFFGMHVQEY GSDCPPPNQR RVYISYLDSV HFFRPKCLRT AVYHEILIGY LEYVKKLGYT TGHIWACPPS EGDD YIFHC HPPDQKIPKP KRLQEWYKKM LDKAVSERIV HDYKDIFKQA TEDRLTSAKE LPYFEGDFWP NVLEESIKEL EQEEE ERKR EENTSNESTD VTKGDSKNAK KKNNKKTSKN KSSLSRGNKK KPGMPNVSND LSQKLYATME KHKEVFFVIR LIAGPA ANS LPPIVDPDPL IPCDLMDGRD AFLTLARDKH LEFSSLRRAQ WSTMCMLVEL HTQSQD UniProtKB: Histone acetyltransferase p300 |
-Macromolecule #2: Cellular tumor antigen p53
| Macromolecule | Name: Cellular tumor antigen p53 / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 29.898908 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: PLSSSVPSQK TYQGSYGFRL GFLHSGTAKS VTCTYSPALN KMFCQLAKTC PVQLWVDSTP PPGTRVRAMA IYKQSQHMTE VVRRCPHHE RCSDSDGLAP PQHLIRVEGN LRVEYLDDRN TFRHSVVVPY EPPEVGSDCT TIHYNYMCNS SCMGGMNRRP I LTIITLED ...String: PLSSSVPSQK TYQGSYGFRL GFLHSGTAKS VTCTYSPALN KMFCQLAKTC PVQLWVDSTP PPGTRVRAMA IYKQSQHMTE VVRRCPHHE RCSDSDGLAP PQHLIRVEGN LRVEYLDDRN TFRHSVVVPY EPPEVGSDCT TIHYNYMCNS SCMGGMNRRP I LTIITLED SSGNLLGRNS FEVRVCACPG RDRRTEEENL RKKGEPHHEL PPGSTKRALP NNTSSSPQPK KKPLDGEYFT LQ IRGRERF EMFRELNEAL ELKDAQAG UniProtKB: Cellular tumor antigen p53 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 279 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Image recording | Film or detector model: FEI EAGLE (4k x 4k) / Average exposure time: 1.0 sec. / Average electron dose: 15.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 150.0 µm / Calibrated defocus max: 4.5 µm / Calibrated defocus min: 1.7 µm / Calibrated magnification: 78894 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 4.5 µm / Nominal defocus min: 1.7 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder model: GATAN 910 MULTI-SPECIMEN SINGLE TILT CRYO TRANSFER HOLDER Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A / Chain - Residue range: 1046-1664 / Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Output model |  PDB-5xzc: |
 Movie
Movie Controller
Controller









































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)