+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6751 | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Coxsackievirus A6 (CVA6) virus A-particle | |||||||||||||||||||||
 Map data Map data | None | |||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | Coxsackievirus A6 / A-particle / icosahedral / VIRUS | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / ribonucleoside triphosphate phosphatase activity / nucleoside-triphosphate phosphatase / channel activity ...symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / ribonucleoside triphosphate phosphatase activity / nucleoside-triphosphate phosphatase / channel activity / monoatomic ion transmembrane transport / DNA replication / RNA helicase activity / endocytosis involved in viral entry into host cell / symbiont-mediated activation of host autophagy / RNA-directed RNA polymerase / cysteine-type endopeptidase activity / viral RNA genome replication / RNA-directed RNA polymerase activity / DNA-templated transcription / virion attachment to host cell / host cell nucleus / structural molecule activity / proteolysis / RNA binding / zinc ion binding / ATP binding Similarity search - Function | |||||||||||||||||||||
| Biological species |  Coxsackievirus A6 Coxsackievirus A6 | |||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||||||||||||||
 Authors Authors | Zheng QB / He MZ | |||||||||||||||||||||
| Funding support |  China, China,  United States, 6 items United States, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2017 Journal: Nat Commun / Year: 2017Title: Atomic structures of Coxsackievirus A6 and its complex with a neutralizing antibody. Authors: Longfa Xu / Qingbing Zheng / Shaowei Li / Maozhou He / Yangtao Wu / Yongchao Li / Rui Zhu / Hai Yu / Qiyang Hong / Jie Jiang / Zizhen Li / Shuxuan Li / Huan Zhao / Lisheng Yang / Wangheng ...Authors: Longfa Xu / Qingbing Zheng / Shaowei Li / Maozhou He / Yangtao Wu / Yongchao Li / Rui Zhu / Hai Yu / Qiyang Hong / Jie Jiang / Zizhen Li / Shuxuan Li / Huan Zhao / Lisheng Yang / Wangheng Hou / Wei Wang / Xiangzhong Ye / Jun Zhang / Timothy S Baker / Tong Cheng / Z Hong Zhou / Xiaodong Yan / Ningshao Xia /   Abstract: Coxsackievirus A6 (CVA6) has recently emerged as a major cause of hand, foot and mouth disease in children worldwide but no vaccine is available against CVA6 infections. Here, we demonstrate the ...Coxsackievirus A6 (CVA6) has recently emerged as a major cause of hand, foot and mouth disease in children worldwide but no vaccine is available against CVA6 infections. Here, we demonstrate the isolation of two forms of stable CVA6 particles-procapsid and A-particle-with excellent biochemical stability and natural antigenicity to serve as vaccine candidates. Despite the presence (in A-particle) or absence (in procapsid) of capsid-RNA interactions, the two CVA6 particles have essentially identical atomic capsid structures resembling the uncoating intermediates of other enteroviruses. Our near-atomic resolution structure of CVA6 A-particle complexed with a neutralizing antibody maps an immune-dominant neutralizing epitope to the surface loops of VP1. The structure-guided cell-based inhibition studies further demonstrate that these loops could serve as excellent targets for designing anti-CVA6 vaccines.Coxsackievirus A6 (CVA6) causes hand, foot and mouth disease in children. Here the authors present the CVA6 procapsid and A-particle cryo-EM structures and identify an immune-dominant neutralizing epitope, which can be exploited for vaccine development. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6751.map.gz emd_6751.map.gz | 246.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6751-v30.xml emd-6751-v30.xml emd-6751.xml emd-6751.xml | 18.3 KB 18.3 KB | Display Display |  EMDB header EMDB header |
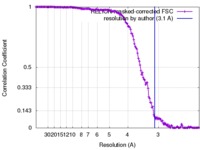
| FSC (resolution estimation) |  emd_6751_fsc.xml emd_6751_fsc.xml | 14.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_6751.png emd_6751.png | 316.7 KB | ||
| Filedesc metadata |  emd-6751.cif.gz emd-6751.cif.gz | 6.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6751 http://ftp.pdbj.org/pub/emdb/structures/EMD-6751 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6751 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6751 | HTTPS FTP |
-Related structure data
| Related structure data |  5xs4MC  6752C  6757C  5xs5C  5xs7C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6751.map.gz / Format: CCP4 / Size: 262.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6751.map.gz / Format: CCP4 / Size: 262.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.128 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Coxsackievirus A6
| Entire | Name:  Coxsackievirus A6 Coxsackievirus A6 |
|---|---|
| Components |
|
-Supramolecule #1: Coxsackievirus A6
| Supramolecule | Name: Coxsackievirus A6 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 86107 / Sci species name: Coxsackievirus A6 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Virus shell | Shell ID: 1 / Name: Coxsackievirus A6 A-particle capsid / T number (triangulation number): 1 |
-Macromolecule #1: Genome polyprotein
| Macromolecule | Name: Genome polyprotein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Coxsackievirus A6 / Organ: Homo sapiens, human / Tissue: muscle Coxsackievirus A6 / Organ: Homo sapiens, human / Tissue: muscle |
| Molecular weight | Theoretical: 33.467273 KDa |
| Sequence | String: NDPIANAVES AVSALADTTI SRVTAANTTA STHSLGTGRV PALQAAETGA SSNASDENLV ETRCVMNRNG VNEASVEHFY SRAGLVGVV EVKDSGTSLD GYTVWPIDVM GFVQQRRKLE LSTYMRFDAE FTFVSNLSNS TTPGMLLQYM YVPPGAPKPD G RKSYQWQT ...String: NDPIANAVES AVSALADTTI SRVTAANTTA STHSLGTGRV PALQAAETGA SSNASDENLV ETRCVMNRNG VNEASVEHFY SRAGLVGVV EVKDSGTSLD GYTVWPIDVM GFVQQRRKLE LSTYMRFDAE FTFVSNLSNS TTPGMLLQYM YVPPGAPKPD G RKSYQWQT ATNPSVFAKL SDPPPQVSVP FMSPATAYQW FYDGYPTFGE HKQATNLQYG QCPNNMMGHF AIRTVSESTT GK NVHVRVY MRIKHVRAWV PRPLRSQAYM VKNYPTYSQT ITNTATDRAS ITTTDYEGGV PASPQRTS UniProtKB: Genome polyprotein |
-Macromolecule #2: Genome polyprotein
| Macromolecule | Name: Genome polyprotein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Coxsackievirus A6 / Organ: homo sapiens, human / Tissue: muscle Coxsackievirus A6 / Organ: homo sapiens, human / Tissue: muscle |
| Molecular weight | Theoretical: 28.095559 KDa |
| Sequence | String: SPSVEACGYS DRVAQLTVGN STITTQEAAN IVLSYGEWPE YCPSTDATAV DKPTRPDVSV NRFYTLSTKS WKTESTGWYW KFPDVLNDT GVFGQNAQFH YLYRSGFCMH VQCNASKFHQ GALLVAAIPE FVIAASSPAT KPNGRGLYPD FAHTNPGKDG Q EFRDPYVL ...String: SPSVEACGYS DRVAQLTVGN STITTQEAAN IVLSYGEWPE YCPSTDATAV DKPTRPDVSV NRFYTLSTKS WKTESTGWYW KFPDVLNDT GVFGQNAQFH YLYRSGFCMH VQCNASKFHQ GALLVAAIPE FVIAASSPAT KPNGRGLYPD FAHTNPGKDG Q EFRDPYVL DAGIPLSQAL VFPHQWINLR TNNCATIIMP YVNALPFDSA LNHSNFGLVV IPISPLKYCN GATTEVPVTL TI APLNSEF SGLRQAIKQ UniProtKB: Genome polyprotein |
-Macromolecule #3: Genome polyprotein
| Macromolecule | Name: Genome polyprotein / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Coxsackievirus A6 / Organ: homo sapiens, human / Tissue: muscle Coxsackievirus A6 / Organ: homo sapiens, human / Tissue: muscle |
| Molecular weight | Theoretical: 26.274836 KDa |
| Sequence | String: GFPTELKPGT NQFLTTDDGT SPPILPGFEP TPLIHIPGEF TSLLDLCQIE TILEVNNTTG TIGVSRLLIP VRAQNNVDQL CASFQVDPG RNGPWQSTMV GQICRYYTQW SGSLKVTFMF TGSFMATGKM LIAYTPPGSA QPATREAAML GTHIVWDFGL Q SSVTLVIP ...String: GFPTELKPGT NQFLTTDDGT SPPILPGFEP TPLIHIPGEF TSLLDLCQIE TILEVNNTTG TIGVSRLLIP VRAQNNVDQL CASFQVDPG RNGPWQSTMV GQICRYYTQW SGSLKVTFMF TGSFMATGKM LIAYTPPGSA QPATREAAML GTHIVWDFGL Q SSVTLVIP WISNTHFRAV KIGGVYDYYA TGIVTIWYQT NFVVPPDTPT EANIIALGAA QKNFTLKLCK DTDEIQQTAA YQ UniProtKB: Genome polyprotein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.4 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Component - Concentration: 20.0 mmol/L / Component - Name: PBS Details: NaCl 137mmol/L, KCl 2.7mmol/L, Na2HPO4 10mmol/L, KH2PO4 2mmol/L |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 40 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV Details: 3?L of virus sample was applied and blot for 6 seconds before plunging.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Digitization - Frames/image: 1-7 / Number grids imaged: 3 / Number real images: 203 / Average exposure time: 1.0 sec. / Average electron dose: 25.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 4.3 µm / Calibrated defocus min: 1.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.3 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 93000 |
| Sample stage | Specimen holder model: GATAN 626 SINGLE TILT LIQUID NITROGEN CRYO TRANSFER HOLDER Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 163.92 / Target criteria: Correlation coefficient |
|---|---|
| Output model |  PDB-5xs4: |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)