+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | native NMDA receptor-GluN1/N2A/N2B-S1 in the closed state | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | native / NMDA receptor / ionotropic ion channel / excitatory neurotransmitter / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationAssembly and cell surface presentation of NMDA receptors / EPHB-mediated forward signaling / Synaptic adhesion-like molecules / Unblocking of NMDA receptors, glutamate binding and activation / neurotransmitter receptor transport, plasma membrane to endosome / RAF/MAP kinase cascade / receptor recycling / sensory organ development / directional locomotion / regulation of cAMP/PKA signal transduction ...Assembly and cell surface presentation of NMDA receptors / EPHB-mediated forward signaling / Synaptic adhesion-like molecules / Unblocking of NMDA receptors, glutamate binding and activation / neurotransmitter receptor transport, plasma membrane to endosome / RAF/MAP kinase cascade / receptor recycling / sensory organ development / directional locomotion / regulation of cAMP/PKA signal transduction / pons maturation / positive regulation of Schwann cell migration / regulation of cell communication / sensitization / olfactory learning / serotonin metabolic process / dendritic branch / fear response / conditioned taste aversion / protein localization to postsynaptic membrane / regulation of ARF protein signal transduction / regulation of respiratory gaseous exchange / apical dendrite / transmitter-gated monoatomic ion channel activity / suckling behavior / sleep / interleukin-1 receptor binding / positive regulation of inhibitory postsynaptic potential / propylene metabolic process / response to glycine / dendritic spine organization / locomotion / negative regulation of dendritic spine maintenance / heterocyclic compound binding / neurotransmitter receptor complex / regulation of monoatomic cation transmembrane transport / NMDA glutamate receptor activity / voltage-gated monoatomic cation channel activity / NMDA selective glutamate receptor complex / glutamate binding / glutamate receptor signaling pathway / ligand-gated sodium channel activity / positive regulation of glutamate secretion / transport vesicle membrane / neuromuscular process / regulation of axonogenesis / calcium ion transmembrane import into cytosol / regulation of dendrite morphogenesis / male mating behavior / regulation of synapse assembly / protein heterotetramerization / response to morphine / small molecule binding / glycine binding / receptor clustering / startle response / dopamine metabolic process / cellular response to zinc ion / positive regulation of reactive oxygen species biosynthetic process / response to lithium ion / parallel fiber to Purkinje cell synapse / regulation of MAPK cascade / monoatomic ion channel complex / behavioral response to pain / regulation of postsynaptic membrane potential / monoatomic cation transmembrane transport / action potential / positive regulation of calcium ion transport into cytosol / extracellularly glutamate-gated ion channel activity / modulation of excitatory postsynaptic potential / associative learning / positive regulation of dendritic spine maintenance / positive regulation of protein targeting to membrane / regulation of neuronal synaptic plasticity / monoatomic cation transport / glutamate receptor binding / detection of mechanical stimulus involved in sensory perception of pain / social behavior / ligand-gated monoatomic ion channel activity / positive regulation of synaptic transmission / phosphatase binding / long-term memory / prepulse inhibition / postsynaptic density, intracellular component / behavioral fear response / monoatomic cation channel activity / synaptic cleft / calcium ion homeostasis / glutamate-gated receptor activity / regulation of long-term synaptic depression / positive regulation of synaptic transmission, glutamatergic / D2 dopamine receptor binding / glutamate-gated calcium ion channel activity / presynaptic active zone membrane / cell adhesion molecule binding / neurogenesis / excitatory synapse / ionotropic glutamate receptor signaling pathway / sensory perception of pain / ionotropic glutamate receptor binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.98 Å | |||||||||
 Authors Authors | Yu J / Xu RS / Ge JP | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2026 Journal: Nature / Year: 2026Title: Conformational diversity and fully opening mechanism of native NMDA receptor. Authors: Ruisheng Xu / Qiqi Jiang / Hongwei Xu / Lu Zhang / Xiangzi Hu / Zizhuo Lu / Huaqin Deng / Haolin Xiong / Sensen Zhang / Zhongwen Chen / Yifan Ge / Zhengjiang Zhu / Yaoyang Zhang / Yelin Chen ...Authors: Ruisheng Xu / Qiqi Jiang / Hongwei Xu / Lu Zhang / Xiangzi Hu / Zizhuo Lu / Huaqin Deng / Haolin Xiong / Sensen Zhang / Zhongwen Chen / Yifan Ge / Zhengjiang Zhu / Yaoyang Zhang / Yelin Chen / Jingpeng Ge / Jie Yu /  Abstract: N-methyl-D-aspartate receptors (NMDARs) are glutamate-gated ion channels that mediate excitatory neurotransmission throughout the brain. As obligate heterotetramers, their activation requires the ...N-methyl-D-aspartate receptors (NMDARs) are glutamate-gated ion channels that mediate excitatory neurotransmission throughout the brain. As obligate heterotetramers, their activation requires the binding of both glycine and glutamate. Although recent structural studies have provided insights into endogenous receptors from select brain regions, most previous work has relied on recombinant receptors and engineered constructs, which limits our understanding of native NMDARs across the whole brain. Here we identify and resolve ten distinct native NMDAR assemblies from the whole-brain tissue of female C57BL/6 mice using immunoaffinity purification, single-molecule total internal reflection fluorescence microscopy and cryo-electron microscopy. Analyses of the GluN1-GluN2A(S1), GluN1-GluN2A(S2), GluN1-GluN2A(S3), GluN1-GluN2B, GluN1-GluN2A-GluN2B(S1), GluN1-GluN2A-GluN2B(S2), GluN1-GluN2A-GluNX(S1), GluN1-GluN2A-GluNX(S2), GluN1-GluN2B-GluNX and GluN1-GluNX structures reveal that GluN2A is the most prevalent subunit across assemblies. Moreover, the substantial conformational flexibility observed in the GluN2A amino-terminal domain may explain its fast kinetics and dominant role in gating. Dynamic movements of S-ketamine were also captured at the channel vestibule, as was pore dilation in both the GluN1 and GluN2B subunits of a native GluN1-GluN2B receptor. The latter observation represents a previously unknown fully open state of NMDAR. Our large collection of heterogeneous NMDAR structures from whole brain reveals previously unrecognized properties of conformational diversity and channel dilation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_64361.map.gz emd_64361.map.gz | 227.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-64361-v30.xml emd-64361-v30.xml emd-64361.xml emd-64361.xml | 21.6 KB 21.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_64361.png emd_64361.png | 37 KB | ||
| Filedesc metadata |  emd-64361.cif.gz emd-64361.cif.gz | 7.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-64361 http://ftp.pdbj.org/pub/emdb/structures/EMD-64361 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-64361 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-64361 | HTTPS FTP |
-Related structure data
| Related structure data |  9unrMC  9un2C  9un3C  9unjC  9unkC  9unmC  9unnC  9unoC  9unpC  9unqC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_64361.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_64361.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.055 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : native NMDA receptor-GluN1/N2A/N2B-S1 in the closed state
| Entire | Name: native NMDA receptor-GluN1/N2A/N2B-S1 in the closed state |
|---|---|
| Components |
|
-Supramolecule #1: native NMDA receptor-GluN1/N2A/N2B-S1 in the closed state
| Supramolecule | Name: native NMDA receptor-GluN1/N2A/N2B-S1 in the closed state type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Glutamate receptor ionotropic, NMDA 1
| Macromolecule | Name: Glutamate receptor ionotropic, NMDA 1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 92.154289 KDa |
| Sequence | String: KIVNIGAVLS TRKHEQMFRE AVNQANKRHG SWKIQLNATS VTHKPNAIQM ALSVCEDLIS SQVYAILVSH PPTPNDHFTP TPVSYTAGF YRIPVLGLTT RMSIYSDKSI HLSFLRTVPP YSHQSSVWFE MMRVYNWNHI ILLVSDDHEG RAAQKRLETL L EERESKAE ...String: KIVNIGAVLS TRKHEQMFRE AVNQANKRHG SWKIQLNATS VTHKPNAIQM ALSVCEDLIS SQVYAILVSH PPTPNDHFTP TPVSYTAGF YRIPVLGLTT RMSIYSDKSI HLSFLRTVPP YSHQSSVWFE MMRVYNWNHI ILLVSDDHEG RAAQKRLETL L EERESKAE KVLQFDPGTK NVTALLMEAR DLEARVIILS ASEDDAATVY RAAAMLNMTG SGYVWLVGER EISGNALRYA PD GIIGLQL INGKNESAHI SDAVGVVAQA VHELLEKENI TDPPRGCVGN TNIWKTGPLF KRVLMSSKYA DGVTGRVEFN EDG DRKFAN YSIMNLQNRK LVQVGIYNGT HVIPNDRKII WPGGETEKPR GYQMSTRLKI VTIHQEPFVY VKPTMSDGTC KEEF TVNGD PVKKVICTGP NDTSPGSPRH TVPQCCYGFC VDLLIKLART MNFTYEVHLV ADGKFGTQER VNNSNKKEWN GMMGE LLSG QADMIVAPLT INNERAQYIE FSKPFKYQGL TILVKKEIPR STLDSFMQPF QSTLWLLVGL SVHVVAVMLY LLDRFS PFG RFKVNSEEEE EDALTLSSAM WFSWGVLLNS GIGEGAPRSF SARILGMVWA GFAMIIVASY TANLAAFLVL DRPEERI TG INDPRLRNPS DKFIYATVKQ SSVDIYFRRQ VELSTMYRHM EKHNYESAAE AIQAVRDNKL HAFIWDSAVL EFEASQKC D LVTTGELFFR SGFGIGMRKD SPWKQNVSLS ILKSHENGFM EDLDKTWVRY QECDSRSNAP ATLTFENMAG VFMLVAGGI VAGIFLIFIE IAYKRHKDA UniProtKB: Glutamate receptor ionotropic, NMDA 1 |
-Macromolecule #2: Glutamate receptor ionotropic, NMDA 2A
| Macromolecule | Name: Glutamate receptor ionotropic, NMDA 2A / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 90.779812 KDa |
| Sequence | String: ALNIAVLLGH SHDVTERELR NLWGPEQATG LPLDVNVVAL LMNRTDPKSL ITHVCDLMSG ARIHGLVFGD DTDQEAVAQM LDFISSQTF IPILGIHGGA SMIMADKDPT STFFQFGASI QQQATVMLKI MQDYDWHVFS LVTTIFPGYR DFISFIKTTV D NSFVGWDM ...String: ALNIAVLLGH SHDVTERELR NLWGPEQATG LPLDVNVVAL LMNRTDPKSL ITHVCDLMSG ARIHGLVFGD DTDQEAVAQM LDFISSQTF IPILGIHGGA SMIMADKDPT STFFQFGASI QQQATVMLKI MQDYDWHVFS LVTTIFPGYR DFISFIKTTV D NSFVGWDM QNVITLDTSF EDAKTQVQLK KIHSSVILLY CSKDEAVLIL SEARSLGLTG YDFFWIVPSL VSGNTELIPK EF PSGLISV SYDDWDYSLE ARVRDGLGIL TTAASSMLEK FSYIPEAKAS CYGQTEKPET PLHTLHQFMV NVTWDGKDLS FTE EGYQVH PRLVVIVLNK DREWEKVGKW ENQTLSLRHA VWPRYKSFSD CEPDDNHLSI VTLEEAPFVI VEDIDPLTET CVRN TVPCR KFVKINNSTN EGMNVKKCCK GFCIDILKKL SRTVKFTYDL YLVTNGKHGK KVNNVWNGMI GEVVYQRAVM AVGSL TINE ERSEVVDFSV PFVETGISVM VSRSNGTVSP SAFLEPFSAS VWVMMFVMLL IVSAIAVFVF EYFSPVGYNR NLAKGK APH GPSFTIGKAI WLLWGLVFNN SVPVQNPKGT TSKIMVSVWA FFAVIFLASY TANLAAFMIQ EEFVDQVTGL SDKKFQR PH DYSPPFRFGT VPNGSTERNI RNNYPYMHQY MTKFNQRGVE DALVSLKTGK LDAFIYDAAV LNYKAGRDEG CKLVTIGS G YIFATTGYGI ALQKGSPWKR QIDLALLQFV GDGEMEELET LWLTGICHNE KNEVMSSQLD IDNMAGVFYM LAAAMALSL ITFIWEHLF UniProtKB: Glutamate receptor ionotropic, NMDA 2A |
-Macromolecule #3: Glutamate receptor ionotropic, NMDA 2B
| Macromolecule | Name: Glutamate receptor ionotropic, NMDA 2B / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 90.6105 KDa |
| Sequence | String: SIGIAVILVG TSDEVAIKDA HEKDDFHHLS VVPRVELVAM NETDPKSIIT RICDLMSDRK IQGVVLADDT DQEAIAQILD FISAQTLTP ILGIHGGSSM IMADKDESSM FFQFGPSIEQ QASVMLNIME EYDWYIFSIV TTYFPGYQDF VNKIRSTIEN S FVGWELEE ...String: SIGIAVILVG TSDEVAIKDA HEKDDFHHLS VVPRVELVAM NETDPKSIIT RICDLMSDRK IQGVVLADDT DQEAIAQILD FISAQTLTP ILGIHGGSSM IMADKDESSM FFQFGPSIEQ QASVMLNIME EYDWYIFSIV TTYFPGYQDF VNKIRSTIEN S FVGWELEE VLLLDMSLDD GDSKIQNQLK KLQSPIILLY CTKEEATYIF EVANSVGLTG YGYTWIVPSL VAGDTDTVPS EF PTGLISV SYDEWDYGLP ARVRDGIAII TTAASDMLSE HSFIPEPKSS CYNTHEKRIY QSNMLNRYLI NVTFEGRNLS FSE DGYQMH PKLVIILLNK ERKWERVGKW KDKSLQMKYY VWPRMCPETE EQEDDHLSIV TLEEAPFVIV ESVDPLSGTC MRNT VPCQK RIISENKTDE EPGYIKKCCK GFCIDILKKI SKSVKFTYDL YLVTNGKHGK KINGTWNGMI GEVVMKRAYM AVGSL TINE ERSEVVDFSV PFIETGISVM VSRSNGTVSP SAFLEPFSAD VWVMMFVMLL IVSAVAVFVF EYFSPVGYNR CLADGR EPG GPSFTIGKAI WLLWGLVFNN SVPVQNPKGT TSKIMVSVWA FFAVIFLASY TANLAAFMIQ EEYVDQVSGL SDKKFQR PN DFSPPFRFGT VPNGSTERNI RNNYAEMHAY MGKFNQRGVD DALLSLKTGK LDAFIYDAAV LNYMAGRDEG CKLVTIGS G KVFASTGYGI AIQKDSGWKR QVDLAILQLF GDGEMEELEA LWLTGICHNE KNEVMSSQLD IDNMAGVFYM LGAAMALSL ITFICEHL UniProtKB: Glutamate receptor ionotropic, NMDA 2B |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 19 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #5: GLYCINE
| Macromolecule | Name: GLYCINE / type: ligand / ID: 5 / Number of copies: 2 / Formula: GLY |
|---|---|
| Molecular weight | Theoretical: 75.067 Da |
| Chemical component information |  ChemComp-GLY: |
-Macromolecule #6: GLUTAMIC ACID
| Macromolecule | Name: GLUTAMIC ACID / type: ligand / ID: 6 / Number of copies: 2 / Formula: GLU |
|---|---|
| Molecular weight | Theoretical: 147.129 Da |
| Chemical component information |  ChemComp-GLU: |
-Macromolecule #7: (2~{S})-2-(2-chlorophenyl)-2-(methylamino)cyclohexan-1-one
| Macromolecule | Name: (2~{S})-2-(2-chlorophenyl)-2-(methylamino)cyclohexan-1-one type: ligand / ID: 7 / Number of copies: 1 / Formula: JC9 |
|---|---|
| Molecular weight | Theoretical: 237.725 Da |
| Chemical component information | 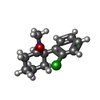 ChemComp-JC9: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 50 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




























































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















