+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | native NMDA receptor-GluN1/N2A/N2B-S2 in the closed state | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | native / NMDA receptor / inotropic ion channel / excitatory neurotransmitter / MEMBRANE / open state / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationAssembly and cell surface presentation of NMDA receptors / EPHB-mediated forward signaling / Synaptic adhesion-like molecules / Unblocking of NMDA receptors, glutamate binding and activation / neurotransmitter receptor transport, plasma membrane to endosome / RAF/MAP kinase cascade / receptor recycling / sensory organ development / directional locomotion / regulation of cAMP/PKA signal transduction ...Assembly and cell surface presentation of NMDA receptors / EPHB-mediated forward signaling / Synaptic adhesion-like molecules / Unblocking of NMDA receptors, glutamate binding and activation / neurotransmitter receptor transport, plasma membrane to endosome / RAF/MAP kinase cascade / receptor recycling / sensory organ development / directional locomotion / regulation of cAMP/PKA signal transduction / pons maturation / positive regulation of Schwann cell migration / regulation of cell communication / sensitization / olfactory learning / serotonin metabolic process / dendritic branch / fear response / conditioned taste aversion / protein localization to postsynaptic membrane / regulation of ARF protein signal transduction / regulation of respiratory gaseous exchange / apical dendrite / transmitter-gated monoatomic ion channel activity / suckling behavior / sleep / interleukin-1 receptor binding / positive regulation of inhibitory postsynaptic potential / propylene metabolic process / response to glycine / dendritic spine organization / locomotion / negative regulation of dendritic spine maintenance / heterocyclic compound binding / neurotransmitter receptor complex / regulation of monoatomic cation transmembrane transport / NMDA glutamate receptor activity / voltage-gated monoatomic cation channel activity / NMDA selective glutamate receptor complex / glutamate binding / glutamate receptor signaling pathway / ligand-gated sodium channel activity / positive regulation of glutamate secretion / transport vesicle membrane / neuromuscular process / regulation of axonogenesis / calcium ion transmembrane import into cytosol / regulation of dendrite morphogenesis / male mating behavior / regulation of synapse assembly / protein heterotetramerization / response to morphine / small molecule binding / glycine binding / receptor clustering / startle response / dopamine metabolic process / cellular response to zinc ion / positive regulation of reactive oxygen species biosynthetic process / response to lithium ion / parallel fiber to Purkinje cell synapse / regulation of MAPK cascade / monoatomic ion channel complex / behavioral response to pain / regulation of postsynaptic membrane potential / monoatomic cation transmembrane transport / action potential / positive regulation of calcium ion transport into cytosol / extracellularly glutamate-gated ion channel activity / modulation of excitatory postsynaptic potential / associative learning / positive regulation of dendritic spine maintenance / positive regulation of protein targeting to membrane / regulation of neuronal synaptic plasticity / monoatomic cation transport / glutamate receptor binding / detection of mechanical stimulus involved in sensory perception of pain / social behavior / ligand-gated monoatomic ion channel activity / positive regulation of synaptic transmission / phosphatase binding / long-term memory / prepulse inhibition / postsynaptic density, intracellular component / behavioral fear response / monoatomic cation channel activity / synaptic cleft / calcium ion homeostasis / glutamate-gated receptor activity / regulation of long-term synaptic depression / positive regulation of synaptic transmission, glutamatergic / D2 dopamine receptor binding / glutamate-gated calcium ion channel activity / presynaptic active zone membrane / cell adhesion molecule binding / neurogenesis / excitatory synapse / ionotropic glutamate receptor signaling pathway / sensory perception of pain / ionotropic glutamate receptor binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.27 Å | |||||||||
 Authors Authors | Yu J / Xu RS / Ge JP | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2026 Journal: Nature / Year: 2026Title: Conformational diversity and fully opening mechanism of native NMDA receptor. Authors: Ruisheng Xu / Qiqi Jiang / Hongwei Xu / Lu Zhang / Xiangzi Hu / Zizhuo Lu / Huaqin Deng / Haolin Xiong / Sensen Zhang / Zhongwen Chen / Yifan Ge / Zhengjiang Zhu / Yaoyang Zhang / Yelin Chen ...Authors: Ruisheng Xu / Qiqi Jiang / Hongwei Xu / Lu Zhang / Xiangzi Hu / Zizhuo Lu / Huaqin Deng / Haolin Xiong / Sensen Zhang / Zhongwen Chen / Yifan Ge / Zhengjiang Zhu / Yaoyang Zhang / Yelin Chen / Jingpeng Ge / Jie Yu /  Abstract: N-methyl-D-aspartate receptors (NMDARs) are glutamate-gated ion channels that mediate excitatory neurotransmission throughout the brain. As obligate heterotetramers, their activation requires the ...N-methyl-D-aspartate receptors (NMDARs) are glutamate-gated ion channels that mediate excitatory neurotransmission throughout the brain. As obligate heterotetramers, their activation requires the binding of both glycine and glutamate. Although recent structural studies have provided insights into endogenous receptors from select brain regions, most previous work has relied on recombinant receptors and engineered constructs, which limits our understanding of native NMDARs across the whole brain. Here we identify and resolve ten distinct native NMDAR assemblies from the whole-brain tissue of female C57BL/6 mice using immunoaffinity purification, single-molecule total internal reflection fluorescence microscopy and cryo-electron microscopy. Analyses of the GluN1-GluN2A(S1), GluN1-GluN2A(S2), GluN1-GluN2A(S3), GluN1-GluN2B, GluN1-GluN2A-GluN2B(S1), GluN1-GluN2A-GluN2B(S2), GluN1-GluN2A-GluNX(S1), GluN1-GluN2A-GluNX(S2), GluN1-GluN2B-GluNX and GluN1-GluNX structures reveal that GluN2A is the most prevalent subunit across assemblies. Moreover, the substantial conformational flexibility observed in the GluN2A amino-terminal domain may explain its fast kinetics and dominant role in gating. Dynamic movements of S-ketamine were also captured at the channel vestibule, as was pore dilation in both the GluN1 and GluN2B subunits of a native GluN1-GluN2B receptor. The latter observation represents a previously unknown fully open state of NMDAR. Our large collection of heterogeneous NMDAR structures from whole brain reveals previously unrecognized properties of conformational diversity and channel dilation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_64359.map.gz emd_64359.map.gz | 223.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-64359-v30.xml emd-64359-v30.xml emd-64359.xml emd-64359.xml | 21.7 KB 21.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_64359.png emd_64359.png | 37.1 KB | ||
| Filedesc metadata |  emd-64359.cif.gz emd-64359.cif.gz | 7.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-64359 http://ftp.pdbj.org/pub/emdb/structures/EMD-64359 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-64359 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-64359 | HTTPS FTP |
-Related structure data
| Related structure data |  9unpMC  9un2C  9un3C  9unjC  9unkC  9unmC  9unnC  9unoC  9unqC  9unrC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_64359.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_64359.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.055 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : native NMDA receptor-GluN1/N2A/N2B-S2 in the closed state
| Entire | Name: native NMDA receptor-GluN1/N2A/N2B-S2 in the closed state |
|---|---|
| Components |
|
-Supramolecule #1: native NMDA receptor-GluN1/N2A/N2B-S2 in the closed state
| Supramolecule | Name: native NMDA receptor-GluN1/N2A/N2B-S2 in the closed state type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Glutamate receptor ionotropic, NMDA 1
| Macromolecule | Name: Glutamate receptor ionotropic, NMDA 1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 92.408602 KDa |
| Sequence | String: PKIVNIGAVL STRKHEQMFR EAVNQANKRH GSWKIQLNAT SVTHKPNAIQ MALSVCEDLI SSQVYAILVS HPPTPNDHFT PTPVSYTAG FYRIPVLGLT TRMSIYSDKS IHLSFLRTVP PYSHQSSVWF EMMRVYNWNH IILLVSDDHE GRAAQKRLET L LEERESKA ...String: PKIVNIGAVL STRKHEQMFR EAVNQANKRH GSWKIQLNAT SVTHKPNAIQ MALSVCEDLI SSQVYAILVS HPPTPNDHFT PTPVSYTAG FYRIPVLGLT TRMSIYSDKS IHLSFLRTVP PYSHQSSVWF EMMRVYNWNH IILLVSDDHE GRAAQKRLET L LEERESKA EKVLQFDPGT KNVTALLMEA RDLEARVIIL SASEDDAATV YRAAAMLNMT GSGYVWLVGE REISGNALRY AP DGIIGLQ LINGKNESAH ISDAVGVVAQ AVHELLEKEN ITDPPRGCVG NTNIWKTGPL FKRVLMSSKY ADGVTGRVEF NED GDRKFA NYSIMNLQNR KLVQVGIYNG THVIPNDRKI IWPGGETEKP RGYQMSTRLK IVTIHQEPFV YVKPTMSDGT CKEE FTVNG DPVKKVICTG PNDTSPGSPR HTVPQCCYGF CVDLLIKLAR TMNFTYEVHL VADGKFGTQE RVNNSNKKEW NGMMG ELLS GQADMIVAPL TINNERAQYI EFSKPFKYQG LTILVKKEIP RSTLDSFMQP FQSTLWLLVG LSVHVVAVML YLLDRF SPF GRFKVNSEEE EEDALTLSSA MWFSWGVLLN SGIGEGAPRS FSARILGMVW AGFAMIIVAS YTANLAAFLV LDRPEER IT GINDPRLRNP SDKFIYATVK QSSVDIYFRR QVELSTMYRH MEKHNYESAA EAIQAVRDNK LHAFIWDSAV LEFEASQK C DLVTTGELFF RSGFGIGMRK DSPWKQNVSL SILKSHENGF MEDLDKTWVR YQECDSRSNA PATLTFENMA GVFMLVAGG IVAGIFLIFI EIAYKRHKDA R UniProtKB: Glutamate receptor ionotropic, NMDA 1 |
-Macromolecule #2: Glutamate receptor ionotropic, NMDA 2A
| Macromolecule | Name: Glutamate receptor ionotropic, NMDA 2A / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 90.33525 KDa |
| Sequence | String: NIAVLLGHSH DVTERELRNL WGPEQATGLP LDVNVVALLM NRTDPKSLIT HVCDLMSGAR IHGLVFGDDT DQEAVAQMLD FISSQTFIP ILGIHGGASM IMADKDPTST FFQFGASIQQ QATVMLKIMQ DYDWHVFSLV TTIFPGYRDF ISFIKTTVDN S FVGWDMQN ...String: NIAVLLGHSH DVTERELRNL WGPEQATGLP LDVNVVALLM NRTDPKSLIT HVCDLMSGAR IHGLVFGDDT DQEAVAQMLD FISSQTFIP ILGIHGGASM IMADKDPTST FFQFGASIQQ QATVMLKIMQ DYDWHVFSLV TTIFPGYRDF ISFIKTTVDN S FVGWDMQN VITLDTSFED AKTQVQLKKI HSSVILLYCS KDEAVLILSE ARSLGLTGYD FFWIVPSLVS GNTELIPKEF PS GLISVSY DDWDYSLEAR VRDGLGILTT AASSMLEKFS YIPEAKASCY GQTEKPETPL HTLHQFMVNV TWDGKDLSFT EEG YQVHPR LVVIVLNKDR EWEKVGKWEN QTLSLRHAVW PRYKSFSDCE PDDNHLSIVT LEEAPFVIVE DIDPLTETCV RNTV PCRKF VKINNSTNEG MNVKKCCKGF CIDILKKLSR TVKFTYDLYL VTNGKHGKKV NNVWNGMIGE VVYQRAVMAV GSLTI NEER SEVVDFSVPF VETGISVMVS RSNGTVSPSA FLEPFSASVW VMMFVMLLIV SAIAVFVFEY FSPVGYNRNL AKGKAP HGP SFTIGKAIWL LWGLVFNNSV PVQNPKGTTS KIMVSVWAFF AVIFLASYTA NLAAFMIQEE FVDQVTGLSD KKFQRPH DY SPPFRFGTVP NGSTERNIRN NYPYMHQYMT KFNQRGVEDA LVSLKTGKLD AFIYDAAVLN YKAGRDEGCK LVTIGSGY I FATTGYGIAL QKGSPWKRQI DLALLQFVGD GEMEELETLW LTGICHNEKN EVMSSQLDID NMAGVFYMLA AAMALSLIT FIWEH UniProtKB: Glutamate receptor ionotropic, NMDA 2A |
-Macromolecule #3: Glutamate receptor ionotropic, NMDA 2B
| Macromolecule | Name: Glutamate receptor ionotropic, NMDA 2B / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 90.925859 KDa |
| Sequence | String: APSIGIAVIL VGTSDEVAIK DAHEKDDFHH LSVVPRVELV AMNETDPKSI ITRICDLMSD RKIQGVVLAD DTDQEAIAQI LDFISAQTL TPILGIHGGS SMIMADKDES SMFFQFGPSI EQQASVMLNI MEEYDWYIFS IVTTYFPGYQ DFVNKIRSTI E NSFVGWEL ...String: APSIGIAVIL VGTSDEVAIK DAHEKDDFHH LSVVPRVELV AMNETDPKSI ITRICDLMSD RKIQGVVLAD DTDQEAIAQI LDFISAQTL TPILGIHGGS SMIMADKDES SMFFQFGPSI EQQASVMLNI MEEYDWYIFS IVTTYFPGYQ DFVNKIRSTI E NSFVGWEL EEVLLLDMSL DDGDSKIQNQ LKKLQSPIIL LYCTKEEATY IFEVANSVGL TGYGYTWIVP SLVAGDTDTV PS EFPTGLI SVSYDEWDYG LPARVRDGIA IITTAASDML SEHSFIPEPK SSCYNTHEKR IYQSNMLNRY LINVTFEGRN LSF SEDGYQ MHPKLVIILL NKERKWERVG KWKDKSLQMK YYVWPRMCPE TEEQEDDHLS IVTLEEAPFV IVESVDPLSG TCMR NTVPC QKRIISENKT DEEPGYIKKC CKGFCIDILK KISKSVKFTY DLYLVTNGKH GKKINGTWNG MIGEVVMKRA YMAVG SLTI NEERSEVVDF SVPFIETGIS VMVSRSNGTV SPSAFLEPFS ADVWVMMFVM LLIVSAVAVF VFEYFSPVGY NRCLAD GRE PGGPSFTIGK AIWLLWGLVF NNSVPVQNPK GTTSKIMVSV WAFFAVIFLA SYTANLAAFM IQEEYVDQVS GLSDKKF QR PNDFSPPFRF GTVPNGSTER NIRNNYAEMH AYMGKFNQRG VDDALLSLKT GKLDAFIYDA AVLNYMAGRD EGCKLVTI G SGKVFASTGY GIAIQKDSGW KRQVDLAILQ LFGDGEMEEL EALWLTGICH NEKNEVMSSQ LDIDNMAGVF YMLGAAMAL SLITFICEHL F UniProtKB: Glutamate receptor ionotropic, NMDA 2B |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 16 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #5: GLYCINE
| Macromolecule | Name: GLYCINE / type: ligand / ID: 5 / Number of copies: 2 / Formula: GLY |
|---|---|
| Molecular weight | Theoretical: 75.067 Da |
| Chemical component information |  ChemComp-GLY: |
-Macromolecule #6: GLUTAMIC ACID
| Macromolecule | Name: GLUTAMIC ACID / type: ligand / ID: 6 / Number of copies: 2 / Formula: GLU |
|---|---|
| Molecular weight | Theoretical: 147.129 Da |
| Chemical component information |  ChemComp-GLU: |
-Macromolecule #7: (2~{S})-2-(2-chlorophenyl)-2-(methylamino)cyclohexan-1-one
| Macromolecule | Name: (2~{S})-2-(2-chlorophenyl)-2-(methylamino)cyclohexan-1-one type: ligand / ID: 7 / Number of copies: 1 / Formula: JC9 |
|---|---|
| Molecular weight | Theoretical: 237.725 Da |
| Chemical component information | 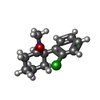 ChemComp-JC9: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




























































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















