[English] 日本語
 Yorodumi
Yorodumi- EMDB-6170: Electron cryo-microscopy of Ltn1, Rqc2 bound to yeast 60S ribosome -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6170 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Electron cryo-microscopy of Ltn1, Rqc2 bound to yeast 60S ribosome | |||||||||
 Map data Map data | 3D class of Ltn1, Rqc2 bound to 60S | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ribosome quality control complex / RQC / eukaryotic ribosome rescue / stalled nascent chain | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 8.2 Å | |||||||||
 Authors Authors | Shen PS / Park J / Qin Y / Li X / Parsawar K / Larson M / Cox J / Cheng Y / Lambowitz AM / Weissman JS ...Shen PS / Park J / Qin Y / Li X / Parsawar K / Larson M / Cox J / Cheng Y / Lambowitz AM / Weissman JS / Brandman O / Frost A | |||||||||
 Citation Citation |  Journal: Science / Year: 2015 Journal: Science / Year: 2015Title: Protein synthesis. Rqc2p and 60S ribosomal subunits mediate mRNA-independent elongation of nascent chains. Authors: Peter S Shen / Joseph Park / Yidan Qin / Xueming Li / Krishna Parsawar / Matthew H Larson / James Cox / Yifan Cheng / Alan M Lambowitz / Jonathan S Weissman / Onn Brandman / Adam Frost /  Abstract: In Eukarya, stalled translation induces 40S dissociation and recruitment of the ribosome quality control complex (RQC) to the 60S subunit, which mediates nascent chain degradation. Here we report ...In Eukarya, stalled translation induces 40S dissociation and recruitment of the ribosome quality control complex (RQC) to the 60S subunit, which mediates nascent chain degradation. Here we report cryo-electron microscopy structures revealing that the RQC components Rqc2p (YPL009C/Tae2) and Ltn1p (YMR247C/Rkr1) bind to the 60S subunit at sites exposed after 40S dissociation, placing the Ltn1p RING (Really Interesting New Gene) domain near the exit channel and Rqc2p over the P-site transfer RNA (tRNA). We further demonstrate that Rqc2p recruits alanine- and threonine-charged tRNA to the A site and directs the elongation of nascent chains independently of mRNA or 40S subunits. Our work uncovers an unexpected mechanism of protein synthesis, in which a protein--not an mRNA--determines tRNA recruitment and the tagging of nascent chains with carboxy-terminal Ala and Thr extensions ("CAT tails"). | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6170.map.gz emd_6170.map.gz | 146.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6170-v30.xml emd-6170-v30.xml emd-6170.xml emd-6170.xml | 10 KB 10 KB | Display Display |  EMDB header EMDB header |
| Images |  400_6170.gif 400_6170.gif 80_6170.gif 80_6170.gif | 62.3 KB 3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6170 http://ftp.pdbj.org/pub/emdb/structures/EMD-6170 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6170 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6170 | HTTPS FTP |
-Related structure data
| Related structure data |  2811C  2812C  6169C  6171C  6172C  6173C  6174C  6175C  6176C  6201C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6170.map.gz / Format: CCP4 / Size: 276 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6170.map.gz / Format: CCP4 / Size: 276 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D class of Ltn1, Rqc2 bound to 60S | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.22 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
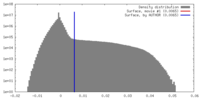
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : RQC particles purified by co-IP of Rqc1-FLAG, eluted with 3xFLAG ...
| Entire | Name: RQC particles purified by co-IP of Rqc1-FLAG, eluted with 3xFLAG peptide |
|---|---|
| Components |
|
-Supramolecule #1000: RQC particles purified by co-IP of Rqc1-FLAG, eluted with 3xFLAG ...
| Supramolecule | Name: RQC particles purified by co-IP of Rqc1-FLAG, eluted with 3xFLAG peptide type: sample / ID: 1000 Details: RQC particles were 3D classified to reveal distinct subclasses containing various RQC components. Number unique components: 1 |
|---|
-Supramolecule #1: 60S ribosome
| Supramolecule | Name: 60S ribosome / type: complex / ID: 1 / Name.synonym: large ribosomal subunit / Recombinant expression: No / Database: NCBI Ribosome-details: ribosome-eukaryote: LSU 60S, LSU RNA 28S, LSU RNA 5.8S, LSU RNA 5S |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 / Details: 100 mM KOAc, 10 mM MgCl2, 25 mM HEPES-KOH |
|---|---|
| Grid | Details: 200 mesh Quantifoil R2/2 grid + lacey carbon grid with ultrathin carbon |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 75 % / Chamber temperature: 90 K / Instrument: FEI VITROBOT MARK III / Method: Blot for 3 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Temperature | Min: 80 K / Max: 90 K / Average: 85 K |
| Specialist optics | Energy filter - Name: Gatan |
| Details | UCSF Image4 on-the-fly motion correction |
| Date | Jul 22, 2013 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 (4k x 4k) / Number real images: 3459 / Average electron dose: 35 e/Å2 / Bits/pixel: 8 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 31000 |
| Sample stage | Specimen holder: LN2 cooled / Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Particles were selected using the semi-automated swarm tool in e2boxer.py of the EMAN2 package. All 2D and 3D processing was performed in RELION. |
|---|---|
| CTF correction | Details: each particle |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 8.2 Å / Resolution method: OTHER / Software - Name: RELION, CTFFIND3 Details: Micrographs were motion-corrected via the UCSFImage4 package. Number images used: 9131 |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)