[English] 日本語
 Yorodumi
Yorodumi- EMDB-6130: Negative stain random conical tilt reconstructions of E. coli rib... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6130 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Negative stain random conical tilt reconstructions of E. coli ribosomal 30S subunit assembly intermediates | |||||||||
 Map data Map data | Group V 30S assembly intermediate missing S3 density from wild-type E. coli (Figure 2F) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ribosome assembly / 30S subunit / Assembly intermediate | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 62.0 Å | |||||||||
 Authors Authors | Sashital DG / Greeman CA / Lyumkis D / Potter CS / Carragher B / Williamson JR | |||||||||
 Citation Citation |  Journal: Elife / Year: 2014 Journal: Elife / Year: 2014Title: A combined quantitative mass spectrometry and electron microscopy analysis of ribosomal 30S subunit assembly in E. coli. Authors: Dipali G Sashital / Candacia A Greeman / Dmitry Lyumkis / Clinton S Potter / Bridget Carragher / James R Williamson /  Abstract: Ribosome assembly is a complex process involving the folding and processing of ribosomal RNAs (rRNAs), concomitant binding of ribosomal proteins (r-proteins), and participation of numerous accessory ...Ribosome assembly is a complex process involving the folding and processing of ribosomal RNAs (rRNAs), concomitant binding of ribosomal proteins (r-proteins), and participation of numerous accessory cofactors. Here, we use a quantitative mass spectrometry/electron microscopy hybrid approach to determine the r-protein composition and conformation of 30S ribosome assembly intermediates in Escherichia coli. The relative timing of assembly of the 3' domain and the formation of the central pseudoknot (PK) structure depends on the presence of the assembly factor RimP. The central PK is unstable in the absence of RimP, resulting in the accumulation of intermediates in which the 3'-domain is unanchored and the 5'-domain is depleted for r-proteins S5 and S12 that contact the central PK. Our results reveal the importance of the cofactor RimP in central PK formation, and introduce a broadly applicable method for characterizing macromolecular assembly in cells. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6130.map.gz emd_6130.map.gz | 20.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6130-v30.xml emd-6130-v30.xml emd-6130.xml emd-6130.xml | 12.2 KB 12.2 KB | Display Display |  EMDB header EMDB header |
| Images |  400_6130.gif 400_6130.gif 80_6130.gif 80_6130.gif | 16.2 KB 2.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6130 http://ftp.pdbj.org/pub/emdb/structures/EMD-6130 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6130 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6130 | HTTPS FTP |
-Related structure data
| Related structure data |  6125C  6126C  6127C  6128C  6129C  6131C  6132C  6133C  6134C  6135C  6136C  6137C  6138C  6139C  6140C  6141C  6142C  6143C 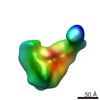 6144C  6145C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6130.map.gz / Format: CCP4 / Size: 41.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6130.map.gz / Format: CCP4 / Size: 41.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Group V 30S assembly intermediate missing S3 density from wild-type E. coli (Figure 2F) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.0504 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
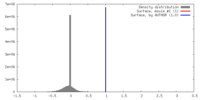
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Group V 30S ribosomal subunit assembly intermediate missing S3 de...
| Entire | Name: Group V 30S ribosomal subunit assembly intermediate missing S3 density from wild-type E. coli |
|---|---|
| Components |
|
-Supramolecule #1000: Group V 30S ribosomal subunit assembly intermediate missing S3 de...
| Supramolecule | Name: Group V 30S ribosomal subunit assembly intermediate missing S3 density from wild-type E. coli type: sample / ID: 1000 Details: Group V particles from heterogeneous sample taken from the center of the 30S sucrose gradient peak Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 820 KDa / Theoretical: 500 KDa Method: Sedimentation and calculation of MW of known components |
-Supramolecule #1: 30S assembly intermediate
| Supramolecule | Name: 30S assembly intermediate / type: complex / ID: 1 / Name.synonym: 30S ribosomal subunit / Details: Particles from center of 30S sucrose gradient peak / Recombinant expression: No / Database: NCBI / Ribosome-details: ribosome-prokaryote: SSU 30S, PSR16s |
|---|---|
| Ref GO | 0: GO:0005840 |
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 820 KDa |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.015 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 20 mM Tris, pH 7.5, 100 mM NH4Cl, 10 mM MgCl2, 0.5 mM EDTA, 6 mM 2-mercaptoethanol |
| Staining | Type: NEGATIVE Details: Negative stain grids were prepared by applying the sample (3 uL) to the grid for 1 min, then blotting from the side to remove excess sample. The grid was washed immediately with 3 uL Buffer ...Details: Negative stain grids were prepared by applying the sample (3 uL) to the grid for 1 min, then blotting from the side to remove excess sample. The grid was washed immediately with 3 uL Buffer A, then blotted from the side. Concurrent with blotting, 3 uL of fresh 2% uranyl formate was applied to the grid, then blotted from the side. This step was repeated twice, then the grid was allowed to dry for at least 10 minutes. |
| Grid | Details: C-flat grids (Protochips) with 2 micron diameter holes coated with a thin (2-5 nm) layer of continuous carbon support, plasma-cleaned for 5s |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI SPIRIT |
|---|---|
| Temperature | Min: 298 K |
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 52,000 times magnification using a live image of the power spectrum. |
| Date | May 27, 2013 |
| Image recording | Category: CCD / Film or detector model: TVIPS TEMCAM-F416 (4k x 4k) / Number real images: 900 / Average electron dose: 20 e/Å2 Details: 450 tilt pairs were collected at -50 and 0 degrees. |
| Tilt angle max | 0 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Calibrated magnification: 52000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 2.25 µm / Nominal defocus min: 0.79 µm / Nominal magnification: 52000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC / Tilt angle min: -50 |
| Experimental equipment |  Model: Tecnai Spirit / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Image tilt pairs were collected (-50 and 0 degrees) and particle tilt pairs were identified and extracted as two separate stacks. The untilted stack was aligned and classified iteratively, and RCT volumes were created for a single class average by applying alignment parameters to the corresponding tilt pairs. |
|---|---|
| CTF correction | Details: Each micrograph |
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 62.0 Å / Resolution method: OTHER / Software - Name: Spider, Appion / Details: RCT reconstruction / Number images used: 248 |
| Final two d classification | Number classes: 1 |
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)