+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1783 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | 30S Ribosome Subunit Assembly Intermediates | |||||||||
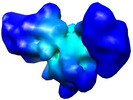
 Map data Map data | 30S Ribosome Assembly Intermediate Group IV a | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ribosome Assembly / 30S Ribosome Subunit / 30S Assembly / Time-resolved electron microscopy / Discovery Single-particle Profiling / 30S Ribosome Subunit Assembly Intermediate | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction | |||||||||
 Authors Authors | Mulder AM / Yoshioka C / Beck A / Bunner A / Milligan RA / Potter CS / Carragher B / Williamson JR | |||||||||
 Citation Citation |  Journal: Science / Year: 2010 Journal: Science / Year: 2010Title: Visualizing ribosome biogenesis: parallel assembly pathways for the 30S subunit. Authors: Anke M Mulder / Craig Yoshioka / Andrea H Beck / Anne E Bunner / Ronald A Milligan / Clinton S Potter / Bridget Carragher / James R Williamson /  Abstract: Ribosomes are self-assembling macromolecular machines that translate DNA into proteins, and an understanding of ribosome biogenesis is central to cellular physiology. Previous studies on the ...Ribosomes are self-assembling macromolecular machines that translate DNA into proteins, and an understanding of ribosome biogenesis is central to cellular physiology. Previous studies on the Escherichia coli 30S subunit suggest that ribosome assembly occurs via multiple parallel pathways rather than through a single rate-limiting step, but little mechanistic information is known about this process. Discovery single-particle profiling (DSP), an application of time-resolved electron microscopy, was used to obtain more than 1 million snapshots of assembling 30S subunits, identify and visualize the structures of 14 assembly intermediates, and monitor the population flux of these intermediates over time. DSP results were integrated with mass spectrometry data to construct the first ribosome-assembly mechanism that incorporates binding dependencies, rate constants, and structural characterization of populated intermediates. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1783.map.gz emd_1783.map.gz | 14.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1783-v30.xml emd-1783-v30.xml emd-1783.xml emd-1783.xml | 7 KB 7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_1783.png emd_1783.png | 103 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1783 http://ftp.pdbj.org/pub/emdb/structures/EMD-1783 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1783 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1783 | HTTPS FTP |
-Related structure data
| Related structure data |  2453C  2454C  2455C  2456C  2457C  2458C  2460C  2461C  2465C  2466C  2467C  2468C  2469C  2470C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1783.map.gz / Format: CCP4 / Size: 100.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1783.map.gz / Format: CCP4 / Size: 100.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 30S Ribosome Assembly Intermediate Group IV a | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.63 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : 30S Reconstitution Reaction
| Entire | Name: 30S Reconstitution Reaction |
|---|---|
| Components |
|
-Supramolecule #1000: 30S Reconstitution Reaction
| Supramolecule | Name: 30S Reconstitution Reaction / type: sample / ID: 1000 / Number unique components: 1 |
|---|---|
| Molecular weight | Theoretical: 789 KDa |
-Supramolecule #1: 30S Ribosome Subunit
| Supramolecule | Name: 30S Ribosome Subunit / type: complex / ID: 1 / Recombinant expression: No / Ribosome-details: ribosome-prokaryote: SSU 30S, PSR16s |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 789 KDa |
-Experimental details
-Structure determination
 Processing Processing | single particle reconstruction |
|---|---|
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Image recording | Number real images: 6635 |
| Electron beam | Acceleration voltage: 120 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder: Single tilt / Specimen holder model: OTHER |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: Ace2 |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Appion, Spider, Eman |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















