[English] 日本語
 Yorodumi
Yorodumi- EMDB-5377: Direct electron detection yields cryo-EM reconstructions at resol... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5377 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Direct electron detection yields cryo-EM reconstructions at resolutions beyond .75 Nyquist frequency | |||||||||
 Map data Map data | Icosahedral reconstruction of bacteriophage epsilon 15 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cryo-EM / Electron cryo-microscopy / Direct detection device / Active pixel sensor / CMOS detector / Nyquist frequency | |||||||||
| Biological species |   Salmonella phage epsilon15 (virus) Salmonella phage epsilon15 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 7.9 Å | |||||||||
 Authors Authors | Bammes BE / Rochat RH / Jakana J / Chen D / Chiu W | |||||||||
 Citation Citation |  Journal: J Struct Biol / Year: 2012 Journal: J Struct Biol / Year: 2012Title: Direct electron detection yields cryo-EM reconstructions at resolutions beyond 3/4 Nyquist frequency. Authors: Benjamin E Bammes / Ryan H Rochat / Joanita Jakana / Dong-Hua Chen / Wah Chiu /  Abstract: One limitation in electron cryo-microscopy (cryo-EM) is the inability to recover high-resolution signal from the image-recording media at the full-resolution limit of the transmission electron ...One limitation in electron cryo-microscopy (cryo-EM) is the inability to recover high-resolution signal from the image-recording media at the full-resolution limit of the transmission electron microscope. Direct electron detection using CMOS-based sensors for digitally recording images has the potential to alleviate this shortcoming. Here, we report a practical performance evaluation of a Direct Detection Device (DDD®) for biological cryo-EM at two different microscope voltages: 200 and 300 kV. Our DDD images of amorphous and graphitized carbon show strong per-pixel contrast with image resolution near the theoretical sampling limit of the data. Single-particle reconstructions of two frozen-hydrated bacteriophages, P22 and ε15, establish that the DDD is capable of recording usable signal for 3D reconstructions at about 4/5 of the Nyquist frequency, which is a vast improvement over the performance of conventional imaging media. We anticipate the unparalleled performance of this digital recording device will dramatically benefit cryo-EM for routine tomographic and single-particle structural determination of biological specimens. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5377.map.gz emd_5377.map.gz | 112.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5377-v30.xml emd-5377-v30.xml emd-5377.xml emd-5377.xml | 10.4 KB 10.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5377_1.jpg emd_5377_1.jpg | 110.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5377 http://ftp.pdbj.org/pub/emdb/structures/EMD-5377 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5377 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5377 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5377.map.gz / Format: CCP4 / Size: 122.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5377.map.gz / Format: CCP4 / Size: 122.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Icosahedral reconstruction of bacteriophage epsilon 15 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.04 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
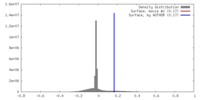
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bacteriophage Epsilon-15
| Entire | Name: Bacteriophage Epsilon-15 |
|---|---|
| Components |
|
-Supramolecule #1000: Bacteriophage Epsilon-15
| Supramolecule | Name: Bacteriophage Epsilon-15 / type: sample / ID: 1000 / Details: Sample was vitrified / Number unique components: 1 |
|---|
-Supramolecule #1: Salmonella phage epsilon15
| Supramolecule | Name: Salmonella phage epsilon15 / type: virus / ID: 1 / Name.synonym: Epsilon 15 / NCBI-ID: 215158 / Sci species name: Salmonella phage epsilon15 / Database: NCBI / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No / Syn species name: Epsilon 15 |
|---|---|
| Host (natural) | Organism:  Salmonella (bacteria) / synonym: BACTERIA(EUBACTERIA) Salmonella (bacteria) / synonym: BACTERIA(EUBACTERIA) |
| Virus shell | Shell ID: 1 / Name: capsid / Diameter: 700 Å / T number (triangulation number): 7 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Details: 400 mesh copper grid |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 170 K / Instrument: OTHER / Details: Vitrification instrument: Vitrobot / Method: Blot for 1.5 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 3200FSC |
|---|---|
| Temperature | Min: 100 K / Max: 100 K / Average: 100 K |
| Alignment procedure | Legacy - Astigmatism: objective lens astigmatism was corrected at 400,000 times magnification |
| Specialist optics | Energy filter - Name: Omega / Energy filter - Lower energy threshold: 15.0 eV / Energy filter - Upper energy threshold: 20.0 eV |
| Date | Mar 6, 2010 |
| Image recording | Category: CCD / Film or detector model: DIRECT ELECTRON DE-12 (4k x 3k) / Digitization - Sampling interval: 6 µm / Number real images: 75 / Average electron dose: 20 e/Å2 Details: Digital images were collected on a CMOS type direct electron detection device DE-12 from Direct Electron |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 20800 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 4.1 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 20000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: JEOL 3200FSC CRYOHOLDER |
- Image processing
Image processing
| Details | Grids were coated in thin carbon before applying sample |
|---|---|
| CTF correction | Details: Each Micrograph |
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 7.9 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: MPSA Details: Resolution was calculated at 7.9 Angstroms by the 0.5 FSC criterion and 6.8 Angstroms by the 0.143 FSC criterion both in comparison with the previously published 4.5 Angstrom resolution reconstruction. Number images used: 1380 |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)