[English] 日本語
 Yorodumi
Yorodumi- EMDB-5221: Nitrogen-Responsive Transcription Factor NrpR form Methanococcus ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5221 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Nitrogen-Responsive Transcription Factor NrpR form Methanococcus maripaludis in an apo form | |||||||||
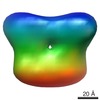
 Map data Map data | Reconstruction volume of apo-NrpR from Methanococcus maripaludis | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Nitrogen assimilation / Nitrogen regulation / Transcriptional regulation | |||||||||
| Function / homology | NrpR regulatory domain Function and homology information Function and homology information | |||||||||
| Biological species |  Methanococcus maripaludis (archaea) Methanococcus maripaludis (archaea) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 25.0 Å | |||||||||
 Authors Authors | Wisedchaisri G / Dranow DM / Lie TJ / Bonanno JB / Patskovsky Y / Ozyurt SA / Sauder JM / Almo SC / Wasserman SR / Burley SK ...Wisedchaisri G / Dranow DM / Lie TJ / Bonanno JB / Patskovsky Y / Ozyurt SA / Sauder JM / Almo SC / Wasserman SR / Burley SK / Leigh JA / Gonen T | |||||||||
 Citation Citation |  Journal: Structure / Year: 2010 Journal: Structure / Year: 2010Title: Structural underpinnings of nitrogen regulation by the prototypical nitrogen-responsive transcriptional factor NrpR. Authors: Goragot Wisedchaisri / David M Dranow / Thomas J Lie / Jeffrey B Bonanno / Yury Patskovsky / Sinem A Ozyurt / J Michael Sauder / Steven C Almo / Stephen R Wasserman / Stephen K Burley / John ...Authors: Goragot Wisedchaisri / David M Dranow / Thomas J Lie / Jeffrey B Bonanno / Yury Patskovsky / Sinem A Ozyurt / J Michael Sauder / Steven C Almo / Stephen R Wasserman / Stephen K Burley / John A Leigh / Tamir Gonen /  Abstract: Plants and microorganisms reduce environmental inorganic nitrogen to ammonium, which then enters various metabolic pathways solely via conversion of 2-oxoglutarate (2OG) to glutamate and glutamine. ...Plants and microorganisms reduce environmental inorganic nitrogen to ammonium, which then enters various metabolic pathways solely via conversion of 2-oxoglutarate (2OG) to glutamate and glutamine. Cellular 2OG concentrations increase during nitrogen starvation. We recently identified a family of 2OG-sensing proteins--the nitrogen regulatory protein NrpR--that bind DNA and repress transcription of nitrogen assimilation genes. We used X-ray crystallography to determine the structure of NrpR regulatory domain. We identified the NrpR 2OG-binding cleft and show that residues predicted to interact directly with 2OG are conserved among diverse classes of 2OG-binding proteins. We show that high levels of 2OG inhibit NrpRs ability to bind DNA. Electron microscopy analyses document that NrpR adopts different quaternary structures in its inhibited 2OG-bound state compared with its active apo state. Our results indicate that upon 2OG release, NrpR repositions its DNA-binding domains correctly for optimal interaction with DNA thereby enabling gene repression. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5221.map.gz emd_5221.map.gz | 247.4 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5221-v30.xml emd-5221-v30.xml emd-5221.xml emd-5221.xml | 10.1 KB 10.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5221_1.jpg emd_5221_1.jpg | 41.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5221 http://ftp.pdbj.org/pub/emdb/structures/EMD-5221 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5221 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5221 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5221.map.gz / Format: CCP4 / Size: 825.2 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5221.map.gz / Format: CCP4 / Size: 825.2 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction volume of apo-NrpR from Methanococcus maripaludis | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.2 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : NrpR from Methanococcus maripaludis
| Entire | Name: NrpR from Methanococcus maripaludis |
|---|---|
| Components |
|
-Supramolecule #1000: NrpR from Methanococcus maripaludis
| Supramolecule | Name: NrpR from Methanococcus maripaludis / type: sample / ID: 1000 / Oligomeric state: Dimer / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 600 KDa / Theoretical: 600 KDa |
-Macromolecule #1: NrpR
| Macromolecule | Name: NrpR / type: protein_or_peptide / ID: 1 / Name.synonym: MMP0607 / Number of copies: 2 / Oligomeric state: Dimer / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  Methanococcus maripaludis (archaea) / Strain: Mm500RC / Location in cell: Cytoplasm Methanococcus maripaludis (archaea) / Strain: Mm500RC / Location in cell: Cytoplasm |
| Molecular weight | Theoretical: 600 KDa |
| Sequence | InterPro: NrpR regulatory domain |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.01 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 100mM Tris HCl pH 7.5, 1M KCl and 5mM glycerol |
| Staining | Type: NEGATIVE Details: A 2 microlitre drop of NrpR was applied to a carbon-coated grid, washed 3 times with milliQ water and stained using 0.075% uranyl formate. |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 12 |
|---|---|
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: NIKON SUPER COOLSCAN 9000 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal magnification: 52000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: OTHER |
- Image processing
Image processing
| CTF correction | Details: Each particle |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 25.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Spider Details: Random conical tilt followed by angular refinement and reconstruction Number images used: 8200 |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | PDBEntryID_givenInChain. Manual docking |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















