+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Endophilin B1 dimer bound to nanodisc center | |||||||||
 Map data Map data | Focused map of an Endophilin B1 dimer bound to the center of a nanodisc | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | BAR / N-BAR / endophilin / membrane / curvature / cardiolipin / MSP2N2 / nanodisc / peripheral membrane protein / APOPTOSIS | |||||||||
| Function / homology |  Function and homology information Function and homology informationlysophosphatidic acid acyltransferase activity / 'de novo' post-translational protein folding / positive regulation of membrane tubulation / phosphatidic acid biosynthetic process / autophagic cell death / protein localization to vacuolar membrane / positive regulation of autophagosome assembly / phosphatidylinositol 3-kinase activator activity / receptor catabolic process / membrane fission ...lysophosphatidic acid acyltransferase activity / 'de novo' post-translational protein folding / positive regulation of membrane tubulation / phosphatidic acid biosynthetic process / autophagic cell death / protein localization to vacuolar membrane / positive regulation of autophagosome assembly / phosphatidylinositol 3-kinase activator activity / receptor catabolic process / membrane fission / membrane organization / : / autophagosome membrane / regulation of macroautophagy / cellular response to glucose starvation / positive regulation of autophagy / fatty acid binding / cellular response to amino acid starvation / regulation of cytokinesis / regulation of protein stability / positive regulation of protein-containing complex assembly / autophagy / cytoplasmic vesicle / midbody / mitochondrial outer membrane / nuclear body / cadherin binding / Golgi membrane / apoptotic process / protein homodimerization activity / protein-containing complex / membrane / identical protein binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
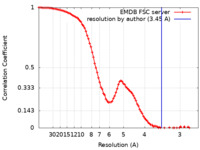
| Method | single particle reconstruction / cryo EM / Resolution: 3.45 Å | |||||||||
 Authors Authors | Thorlacius A / Sundborger-Lunna A | |||||||||
| Funding support |  Sweden, 1 items Sweden, 1 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2025 Journal: Commun Biol / Year: 2025Title: Peripheral membrane protein endophilin B1 probes, perturbs and permeabilizes lipid bilayers. Authors: Arni Thorlacius / Maksim Rulev / Oscar Sundberg / Anna Sundborger-Lunna /  Abstract: Bin/Amphiphysin/Rvs167 (BAR) domain containing proteins are peripheral membrane proteins that regulate intracellular membrane curvature. BAR protein endophilin B1 plays a key role in multiple ...Bin/Amphiphysin/Rvs167 (BAR) domain containing proteins are peripheral membrane proteins that regulate intracellular membrane curvature. BAR protein endophilin B1 plays a key role in multiple cellular processes critical for oncogenesis, including autophagy and apoptosis. Amphipathic regions in endophilin B1 drive membrane association and tubulation through membrane scaffolding. Our understanding of exactly how BAR proteins like endophilin B1 promote highly diverse intracellular membrane remodeling events in the cell is severely limited due to lack of high-resolution structural information. Here we present the highest resolution cryo-EM structure of a BAR protein to date and the first structures of a BAR protein bound to a lipid bicelle. Using neural networks, we can effectively sort particle species of different stoichiometries, revealing the tremendous flexibility of post-membrane binding, pre-polymer BAR dimer organization and membrane deformation. We also show that endophilin B1 efficiently permeabilizes negatively charged liposomes that contain mitochondria-specific lipid cardiolipin and propose a new model for Bax-mediated cell death. #1:  Journal: Biorxiv / Year: 2024 Journal: Biorxiv / Year: 2024Title: Peripheral membrane protein endophilin B1 probes, perturbs and permeabilizes lipid bilayers Authors: Thorlacius A / Rulev M / Sundberg O / Sundborger-Lunna A | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_50984.map.gz emd_50984.map.gz | 32.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-50984-v30.xml emd-50984-v30.xml emd-50984.xml emd-50984.xml | 16.7 KB 16.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_50984_fsc.xml emd_50984_fsc.xml | 11.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_50984.png emd_50984.png | 24.9 KB | ||
| Masks |  emd_50984_msk_1.map emd_50984_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-50984.cif.gz emd-50984.cif.gz | 6.3 KB | ||
| Others |  emd_50984_half_map_1.map.gz emd_50984_half_map_1.map.gz emd_50984_half_map_2.map.gz emd_50984_half_map_2.map.gz | 59.4 MB 59.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-50984 http://ftp.pdbj.org/pub/emdb/structures/EMD-50984 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-50984 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-50984 | HTTPS FTP |
-Related structure data
| Related structure data |  9g2uMC  9g2rC  9g2wC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_50984.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_50984.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Focused map of an Endophilin B1 dimer bound to the center of a nanodisc | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.328 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_50984_msk_1.map emd_50984_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_50984_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_50984_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Endophilin B1 dimers bound to a nanodisc
| Entire | Name: Endophilin B1 dimers bound to a nanodisc |
|---|---|
| Components |
|
-Supramolecule #1: Endophilin B1 dimers bound to a nanodisc
| Supramolecule | Name: Endophilin B1 dimers bound to a nanodisc / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Endophilin-B1
| Macromolecule | Name: Endophilin-B1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.843246 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNIMDFNVKK LAADAGTFLS RAVQFTEEKL GQAEKTELDA HLENLLSKAE CTKIWTEKIM KQTEVLLQPN PNARIEEFVY EKLDRKAPS RINNPELLGQ YMIDAGTEFG PGTAYGNALI KCGETQKRIG TADRELIQTS ALNFLTPLRN FIEGDYKTIA K ERKLLQNK ...String: MNIMDFNVKK LAADAGTFLS RAVQFTEEKL GQAEKTELDA HLENLLSKAE CTKIWTEKIM KQTEVLLQPN PNARIEEFVY EKLDRKAPS RINNPELLGQ YMIDAGTEFG PGTAYGNALI KCGETQKRIG TADRELIQTS ALNFLTPLRN FIEGDYKTIA K ERKLLQNK RLDLDAAKTR LKKAKAAETR NSSEQELRIT QSEFDRQAEI TRLLLEGISS THAHHLRCLN DFVEAQMTYY AQ CYQYMLD LQKQLGSFPS NYLSNNNQTS VTPVPSVLPN AIGSSAMAST SGLVITSPSN LSDLKECSGS RKARVLYDYD AAN STELSL LADEVITVFS VVGMDSDWLM GERGNQKGKV PITYLELLN UniProtKB: Endophilin-B1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.8 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 20 mM Tris-HCl, 100 mM NaCl, 0.5 mM EDTA, pH 7.4 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 130000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A / Chain - Residue range: 11-252 / Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Details | Alphafold model was processed in PHENIX, docked into the density map using UCSF Chimera, refined using all-chain refinement in coot followed by real-space refinement with Servalcat. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
| Output model |  PDB-9g2u: |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)