+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The structure of GluA1/A4 LBD-TMD with 2 TARPs | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | iGluR / CP-AMPA receptors / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationCargo concentration in the ER / Presynaptic depolarization and calcium channel opening / axonal spine / positive regulation of locomotion involved in locomotory behavior / COPII-mediated vesicle transport / positive regulation of membrane potential / eye blink reflex / positive regulation of protein localization to basolateral plasma membrane / cellular response to ammonium ion / response to sucrose ...Cargo concentration in the ER / Presynaptic depolarization and calcium channel opening / axonal spine / positive regulation of locomotion involved in locomotory behavior / COPII-mediated vesicle transport / positive regulation of membrane potential / eye blink reflex / positive regulation of protein localization to basolateral plasma membrane / cellular response to ammonium ion / response to sucrose / myosin V binding / cerebellar mossy fiber / LGI-ADAM interactions / neuron spine / Trafficking of AMPA receptors / postsynaptic neurotransmitter receptor diffusion trapping / proximal dendrite / regulation of AMPA receptor activity / kainate selective glutamate receptor complex / response to arsenic-containing substance / regulation of monoatomic ion transmembrane transport / channel regulator activity / cellular response to L-glutamate / membrane hyperpolarization / ligand-gated calcium channel activity / regulation of synapse structure or activity / cellular response to dsRNA / dendritic spine membrane / nervous system process / beta-2 adrenergic receptor binding / Synaptic adhesion-like molecules / long-term synaptic depression / protein targeting to membrane / cellular response to peptide hormone stimulus / voltage-gated calcium channel complex / cellular response to amine stimulus / peptide hormone receptor binding / response to psychosocial stress / Activation of AMPA receptors / neurotransmitter receptor localization to postsynaptic specialization membrane / response to morphine / perisynaptic space / spinal cord development / neuronal cell body membrane / protein kinase A binding / neuromuscular junction development / Trafficking of GluR2-containing AMPA receptors / negative regulation of smooth muscle cell apoptotic process / response to lithium ion / AMPA glutamate receptor activity / transmission of nerve impulse / immunoglobulin binding / adenylate cyclase binding / behavioral response to pain / asymmetric synapse / AMPA glutamate receptor complex / regulation of receptor recycling / response to electrical stimulus / cellular response to glycine / membrane depolarization / ionotropic glutamate receptor complex / Unblocking of NMDA receptors, glutamate binding and activation / G-protein alpha-subunit binding / conditioned place preference / glutamate receptor binding / positive regulation of synaptic transmission / regulation of postsynaptic membrane neurotransmitter receptor levels / voltage-gated calcium channel activity / long-term memory / postsynaptic density, intracellular component / response to fungicide / neuronal action potential / positive regulation of synaptic transmission, glutamatergic / glutamate-gated receptor activity / cellular response to brain-derived neurotrophic factor stimulus / somatodendritic compartment / glutamate-gated calcium ion channel activity / synapse assembly / presynaptic active zone membrane / excitatory synapse / ionotropic glutamate receptor binding / dendrite membrane / ligand-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential / positive regulation of excitatory postsynaptic potential / hippocampal mossy fiber to CA3 synapse / dendritic shaft / synaptic membrane / calcium channel regulator activity / PDZ domain binding / response to cocaine / neuromuscular junction / cellular response to amino acid stimulus / synaptic transmission, glutamatergic / regulation of membrane potential / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / response to nutrient levels / cerebral cortex development / receptor internalization / regulation of synaptic plasticity / response to calcium ion Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.22 Å | |||||||||
 Authors Authors | Fang CL / Gouaux E | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2025 Journal: Nature / Year: 2025Title: Gating and noelin clustering of native Ca-permeable AMPA receptors. Authors: Chengli Fang / Cathy J Spangler / Jumi Park / Natalie Sheldon / Laurence O Trussell / Eric Gouaux /  Abstract: AMPA-type ionotropic glutamate receptors (AMPARs) are integral to fast excitatory synaptic transmission and have vital roles in synaptic plasticity, motor coordination, learning and memory. Whereas ...AMPA-type ionotropic glutamate receptors (AMPARs) are integral to fast excitatory synaptic transmission and have vital roles in synaptic plasticity, motor coordination, learning and memory. Whereas extensive structural studies have been conducted on recombinant AMPARs and native calcium-impermeable (CI)-AMPARs alongside their auxiliary proteins, the molecular architecture of native calcium-permeable (CP)-AMPARs has remained undefined. Here, to determine the subunit composition, physiological architecture and gating mechanisms of CP-AMPARs, we visualize these receptors, immunoaffinity purified from rat cerebella, and resolve their structures using cryo-electron microscopy (cryo-EM). Our results indicate that the predominant assembly consists of GluA1 and GluA4 subunits, with the GluA4 subunit occupying the B and D positions, and auxiliary subunits, including transmembrane AMPAR regulatory proteins (TARPs) located at the B' and D' positions, and cornichon homologues (CNIHs) or TARPs located at the A' and C' positions. Furthermore, we resolved the structure of the noelin (NOE1)-GluA1-GluA4 complex, in which NOE1 specifically binds to the GluA4 subunit at the B and D positions. Notably, NOE1 stabilizes the amino-terminal domain layer without affecting gating properties of the receptor. NOE1 contributes to AMPAR function by forming dimeric AMPAR assemblies that are likely to engage in extracellular networks, clustering receptors in synaptic environments and modulating receptor responsiveness to synaptic inputs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_49726.map.gz emd_49726.map.gz | 40.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-49726-v30.xml emd-49726-v30.xml emd-49726.xml emd-49726.xml | 22.6 KB 22.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_49726_fsc.xml emd_49726_fsc.xml | 7.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_49726.png emd_49726.png | 93.9 KB | ||
| Filedesc metadata |  emd-49726.cif.gz emd-49726.cif.gz | 6.6 KB | ||
| Others |  emd_49726_additional_1.map.gz emd_49726_additional_1.map.gz emd_49726_half_map_1.map.gz emd_49726_half_map_1.map.gz emd_49726_half_map_2.map.gz emd_49726_half_map_2.map.gz | 21.7 MB 39.6 MB 39.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-49726 http://ftp.pdbj.org/pub/emdb/structures/EMD-49726 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-49726 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-49726 | HTTPS FTP |
-Related structure data
| Related structure data |  9nr9MC  9nr6C  9nr7C  9nr8C  9nraC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_49726.map.gz / Format: CCP4 / Size: 42.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_49726.map.gz / Format: CCP4 / Size: 42.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.88 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_49726_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_49726_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_49726_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CP-AMPA receptors
| Entire | Name: CP-AMPA receptors |
|---|---|
| Components |
|
-Supramolecule #1: CP-AMPA receptors
| Supramolecule | Name: CP-AMPA receptors / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Voltage-dependent calcium channel gamma-2 subunit
| Macromolecule | Name: Voltage-dependent calcium channel gamma-2 subunit / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 22.589975 KDa |
| Sequence | String: DRGVQMLLTT VGAFAAFSLM TIAVGTDYWL YSRGVCKTKS VSENETSKKN EEVMTHSGLW RTCCLEGNFK GLCKQIDHFP EDADYEADT AEYFLRAVRA SSIFPILSVI LLFMGGLCIA ASEFYKTRHN IILSAGIFFV SAGLSNIIGI IVYISANAGD P SKSDSKKN ...String: DRGVQMLLTT VGAFAAFSLM TIAVGTDYWL YSRGVCKTKS VSENETSKKN EEVMTHSGLW RTCCLEGNFK GLCKQIDHFP EDADYEADT AEYFLRAVRA SSIFPILSVI LLFMGGLCIA ASEFYKTRHN IILSAGIFFV SAGLSNIIGI IVYISANAGD P SKSDSKKN SYSYGWSFYF GALSFIIAEM VGVLAVHMFI DRHKQL UniProtKB: Voltage-dependent calcium channel gamma-2 subunit |
-Macromolecule #2: Glutamate receptor 1
| Macromolecule | Name: Glutamate receptor 1 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 47.879855 KDa |
| Sequence | String: RTYIVTTILE DPYVMLKKNA NQFEGNDRYE GYCVELAAEI AKHVGYSYRL EIVSDGKYGA RDPDTKAWNG MVGELVYGRA DVAVAPLTI TLVREEVIDF SKPFMSLGIS IMIKKPQKSK PGVFSFLDPL AYEIWMCIVF AYIGVSVVLF LVSRFSPYEW H SEEFEEGR ...String: RTYIVTTILE DPYVMLKKNA NQFEGNDRYE GYCVELAAEI AKHVGYSYRL EIVSDGKYGA RDPDTKAWNG MVGELVYGRA DVAVAPLTI TLVREEVIDF SKPFMSLGIS IMIKKPQKSK PGVFSFLDPL AYEIWMCIVF AYIGVSVVLF LVSRFSPYEW H SEEFEEGR DQTTSDQSNE FGIFNSLWFS LGAFMQQGCD ISPRSLSGRI VGGVWWFFTL IIISSYTANL AAFLTVERMV SP IESAEDL AKQTEIAYGT LEAGSTKEFF RRSKIAVFEK MWTYMKSAEP SVFVRTTEEG MIRVRKSKGK YAYLLESTMN EYI EQRKPC DTMKVGGNLD SKGYGIATPK GSALRNPVNL AVLKLNEQGL LDKLKNKWWY DKGECGSGGG DSKDKTSALS LSNV AGVFY ILIGGLGLAM LVALIEFCYK SR UniProtKB: Glutamate receptor 1 |
-Macromolecule #3: Isoform 2 of Glutamate receptor 4
| Macromolecule | Name: Isoform 2 of Glutamate receptor 4 / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 47.173348 KDa |
| Sequence | String: VVVTTIMESP YVMYKKNHEM FEGNDKYEGY CVDLASEIAK HIGIKYKIAI VPDGKYGARD ADTKIWNGMV GELVYGKAEI AIAPLTITL VREEVIDFSK PFMSLGISIM IKKPQKSKPG VFSFLDPLAY EIWMCIVFAY IGVSVVLFLV SRFSPYEWHT E EPEDGKEG ...String: VVVTTIMESP YVMYKKNHEM FEGNDKYEGY CVDLASEIAK HIGIKYKIAI VPDGKYGARD ADTKIWNGMV GELVYGKAEI AIAPLTITL VREEVIDFSK PFMSLGISIM IKKPQKSKPG VFSFLDPLAY EIWMCIVFAY IGVSVVLFLV SRFSPYEWHT E EPEDGKEG PSDQPPNEFG IFNSLWFSLG AFMQQGCDIS PRSLSGRIVG GVWWFFTLII ISSYTANLAA FLTVERMVSP IE SAEDLAK QTEIAYGTLD SGSTKEFFRR SKIAVYEKMW TYMRSAEPSV FTRTTAEGVA RVRKSKGKFA FLLESTMNEY TEQ RKPCDT MKVGGNLDSK GYGVATPKGS SLRTPVNLAV LKLSEAGVLD KLKNKWWYDK GECGPKDSGS KDKTSALSLS NVAG VFYIL VGGLGLAMLV ALIEFCYKS UniProtKB: Glutamate receptor 4 |
-Macromolecule #4: {[7-morpholin-4-yl-2,3-dioxo-6-(trifluoromethyl)-3,4-dihydroquino...
| Macromolecule | Name: {[7-morpholin-4-yl-2,3-dioxo-6-(trifluoromethyl)-3,4-dihydroquinoxalin-1(2H)-yl]methyl}phosphonic acid type: ligand / ID: 4 / Number of copies: 4 / Formula: ZK1 |
|---|---|
| Molecular weight | Theoretical: 409.254 Da |
| Chemical component information | 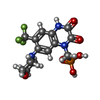 ChemComp-ZK1: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: OTHER / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)













































