+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human TOP3B-TDRD3 core complex in DNA pre-cleavage state | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Topoisomerase / DNA / ISOMERASE-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationDNA topoisomerase III-beta-TDRD3 complex / RNA topoisomerase activity / DNA topoisomerase activity / DNA topoisomerase / DNA topoisomerase type I (single strand cut, ATP-independent) activity / DNA topological change / condensed chromosome / : / chromosome segregation / chromatin organization ...DNA topoisomerase III-beta-TDRD3 complex / RNA topoisomerase activity / DNA topoisomerase activity / DNA topoisomerase / DNA topoisomerase type I (single strand cut, ATP-independent) activity / DNA topological change / condensed chromosome / : / chromosome segregation / chromatin organization / DNA recombination / transcription coactivator activity / DNA repair / chromatin binding / Golgi apparatus / DNA binding / RNA binding / nucleoplasm / nucleus / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.35 Å | |||||||||
 Authors Authors | Yang X / Chen X / Yang W / Pommier Y | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2025 Journal: Nat Commun / Year: 2025Title: Structural insights into human topoisomerase 3β DNA and RNA catalysis and nucleic acid gate dynamics. Authors: Xi Yang / Xuemin Chen / Wei Yang / Yves Pommier /   Abstract: Type IA topoisomerases (TopoIAs) are present in all living organisms. They resolve DNA/RNA catenanes, knots and supercoils by breaking and rejoining single-stranded DNA/RNA segments and allowing the ...Type IA topoisomerases (TopoIAs) are present in all living organisms. They resolve DNA/RNA catenanes, knots and supercoils by breaking and rejoining single-stranded DNA/RNA segments and allowing the passage of another nucleic acid segment through the break. Topoisomerase III-β (TOP3B), the only RNA topoisomerase in metazoans, promotes R-loop disassembly and translation of mRNAs. Defects in TOP3B lead to severe neurological diseases. We present a series of cryo-EM structures of human TOP3B with its cofactor TDRD3 during cleavage and rejoining of DNA or RNA, thus elucidating the roles of divalent metal ions and key enzyme residues in each step of the catalytic cycle. We also obtained the structure of an open-gate configuration that addresses the long-standing question of the strand-passage mechanism. Our studies reveal how TOP3B catalyzes both DNA and RNA relaxation, while TOP3A acts only on DNA. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_45376.map.gz emd_45376.map.gz | 143.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-45376-v30.xml emd-45376-v30.xml emd-45376.xml emd-45376.xml | 15.8 KB 15.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_45376.png emd_45376.png | 41.4 KB | ||
| Filedesc metadata |  emd-45376.cif.gz emd-45376.cif.gz | 6.4 KB | ||
| Others |  emd_45376_half_map_1.map.gz emd_45376_half_map_1.map.gz emd_45376_half_map_2.map.gz emd_45376_half_map_2.map.gz | 154.2 MB 154.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-45376 http://ftp.pdbj.org/pub/emdb/structures/EMD-45376 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-45376 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-45376 | HTTPS FTP |
-Validation report
| Summary document |  emd_45376_validation.pdf.gz emd_45376_validation.pdf.gz | 889.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_45376_full_validation.pdf.gz emd_45376_full_validation.pdf.gz | 889.4 KB | Display | |
| Data in XML |  emd_45376_validation.xml.gz emd_45376_validation.xml.gz | 14.7 KB | Display | |
| Data in CIF |  emd_45376_validation.cif.gz emd_45376_validation.cif.gz | 17.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-45376 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-45376 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-45376 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-45376 | HTTPS FTP |
-Related structure data
| Related structure data |  9c9yMC  9c9wC  9ca0C  9ca1C  9ca4C  9cagC  9cahC  9cajC  9cakC  9calC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_45376.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_45376.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
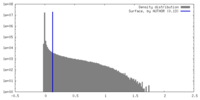
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_45376_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_45376_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : human TOP3B-TDRD3 core complex with DNA
| Entire | Name: human TOP3B-TDRD3 core complex with DNA |
|---|---|
| Components |
|
-Supramolecule #1: human TOP3B-TDRD3 core complex with DNA
| Supramolecule | Name: human TOP3B-TDRD3 core complex with DNA / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: DNA topoisomerase 3-beta-1
| Macromolecule | Name: DNA topoisomerase 3-beta-1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: DNA topoisomerase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 69.021875 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: VMKTVLMVAE KPSLAQSIAK ILSRGSLSSH KGLNGACSVH EYTGTFAGQP VRFKMTSVCG HVMTLDFLGK YNKWDKVDPA ELFSQAPTE KKEANPKLNM VKFLQVEGRG CDYIVLWLDC DKEGENICFE VLDAVLPVMN KAHGGEKTVF RARFSSITDT D ICNAMACL ...String: VMKTVLMVAE KPSLAQSIAK ILSRGSLSSH KGLNGACSVH EYTGTFAGQP VRFKMTSVCG HVMTLDFLGK YNKWDKVDPA ELFSQAPTE KKEANPKLNM VKFLQVEGRG CDYIVLWLDC DKEGENICFE VLDAVLPVMN KAHGGEKTVF RARFSSITDT D ICNAMACL GEPDHNEALS VDARQELDLR IGCAFTRFQT KYFQGKYGDL DSSLISFGPC QTPTLGFCVE RHDKIQSFKP ET YWVLQAK VNTDKDRSLL LDWDRVRVFD REIAQMFLNM TKLEKEAQVE ATSRKEKAKQ RPLALNTVEM LRVASSSLGM GPQ HAMQTA ERLYTQGYIS FPRTETTHYP ENFDLKGSLR QQANHPYWAD TVKRLLAEGI NRPRKGHDAG DHPPITPMKS ATEA ELGGD AWRLYEYITR HFIATVSHDC KYLQSTISFR IGPELFTCSG KTVLSPGFTE VMPWQSVPLE ESLPTCQRGD AFPVG EVKM LEKQTNPPDY LTEAELITLM EKHGIGTDAS IPVHINNICQ RNYVTVESGR RLKPTNLGIV LVHGYYKIDA ELVLPT IRS AVEKQLNLIA QGKADYRQVL GHTLDVFKRK FHYFVDSIAG MDELMEVSF UniProtKB: DNA topoisomerase 3-beta-1 |
-Macromolecule #2: Tudor domain-containing protein 3
| Macromolecule | Name: Tudor domain-containing protein 3 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 17.428951 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAQVAGAALS QAGWYLSDEG IEACTSSPDK VNVNDIILIA LNTDLRTIGK KFLPSDINSG KVEKLEGPCV LQIQKIRNVA APKDNEESQ AAPRMLRLQM TDGHISCTAV EFSYMSKISL NTPPGTKVKL SGIVDIKNGF LLLNDSNTTV LGGEVEHLIE K W UniProtKB: Tudor domain-containing protein 3 |
-Macromolecule #3: DNA (5'-D(P*AP*CP*TP*AP*AP*AP*AP*T)-3')
| Macromolecule | Name: DNA (5'-D(P*AP*CP*TP*AP*AP*AP*AP*T)-3') / type: dna / ID: 3 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 2.418644 KDa |
| Sequence | String: (DA)(DC)(DT)(DA)(DA)(DA)(DA)(DT) |
-Macromolecule #4: MANGANESE (II) ION
| Macromolecule | Name: MANGANESE (II) ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: MN |
|---|---|
| Molecular weight | Theoretical: 54.938 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 46.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.35 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 720245 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)