+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of V. cholerae monomeric DdmD bound with ssDNA | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | DNA defense modules(Ddm) / DdmDE / Vibrio Cholerae / bacteria / anti-plasmid / IMMUNE SYSTEM-DNA complex | |||||||||
| Function / homology | Helicase/UvrB N-terminal domain-containing protein Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
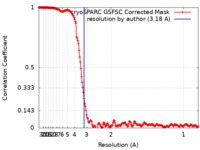
| Method | single particle reconstruction / cryo EM / Resolution: 3.18 Å | |||||||||
 Authors Authors | Shen ZF / Yang XY / Fu TM | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2024 Journal: Cell / Year: 2024Title: DdmDE eliminates plasmid invasion by DNA-guided DNA targeting. Authors: Xiao-Yuan Yang / Zhangfei Shen / Chen Wang / Kotaro Nakanishi / Tian-Min Fu /  Abstract: Horizontal gene transfer is a key driver of bacterial evolution, but it also presents severe risks to bacteria by introducing invasive mobile genetic elements. To counter these threats, bacteria have ...Horizontal gene transfer is a key driver of bacterial evolution, but it also presents severe risks to bacteria by introducing invasive mobile genetic elements. To counter these threats, bacteria have developed various defense systems, including prokaryotic Argonautes (pAgos) and the DNA defense module DdmDE system. Through biochemical analysis, structural determination, and in vivo plasmid clearance assays, we elucidate the assembly and activation mechanisms of DdmDE, which eliminates small, multicopy plasmids. We demonstrate that DdmE, a pAgo-like protein, acts as a catalytically inactive, DNA-guided, DNA-targeting defense module. In the presence of guide DNA, DdmE targets plasmids and recruits a dimeric DdmD, which contains nuclease and helicase domains. Upon binding to DNA substrates, DdmD transitions from an autoinhibited dimer to an active monomer, which then translocates along and cleaves the plasmids. Together, our findings reveal the intricate mechanisms underlying DdmDE-mediated plasmid clearance, offering fundamental insights into bacterial defense systems against plasmid invasions. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_45254.map.gz emd_45254.map.gz | 481.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-45254-v30.xml emd-45254-v30.xml emd-45254.xml emd-45254.xml | 20 KB 20 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_45254_fsc.xml emd_45254_fsc.xml | 19.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_45254.png emd_45254.png | 101.2 KB | ||
| Filedesc metadata |  emd-45254.cif.gz emd-45254.cif.gz | 6.8 KB | ||
| Others |  emd_45254_additional_1.map.gz emd_45254_additional_1.map.gz emd_45254_half_map_1.map.gz emd_45254_half_map_1.map.gz emd_45254_half_map_2.map.gz emd_45254_half_map_2.map.gz | 434.6 MB 473.8 MB 473.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-45254 http://ftp.pdbj.org/pub/emdb/structures/EMD-45254 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-45254 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-45254 | HTTPS FTP |
-Related structure data
| Related structure data |  9c6qMC  9bf1C  9bf5C  9bgkC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_45254.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_45254.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.4495 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_45254_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_45254_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_45254_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Monomeric complex of V. cholerae DdmD bound with ssDNA
| Entire | Name: Monomeric complex of V. cholerae DdmD bound with ssDNA |
|---|---|
| Components |
|
-Supramolecule #1: Monomeric complex of V. cholerae DdmD bound with ssDNA
| Supramolecule | Name: Monomeric complex of V. cholerae DdmD bound with ssDNA type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: ssDNA
| Macromolecule | Name: ssDNA / type: dna / ID: 1 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 3.597367 KDa |
| Sequence | String: (DC)(DT)(DA)(DT)(DT)(DA)(DG)(DC)(DC)(DT) (DC)(DA) |
-Macromolecule #2: Helicase/UvrB N-terminal domain-containing protein
| Macromolecule | Name: Helicase/UvrB N-terminal domain-containing protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 136.155328 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNVSIEEFTH FDFQLVPEPS PLDLVITEPL KNHIEVNGVK SGALLPLPFQ TGIGKTYTAL NFLLQQMLEQ VRSELKEENT GKKSKRLLY YVTDSVDNVV SAKADLLKLI EKQTVKGEPR FTLEQQEYLK AQIVHLPNQS EQLLQCSDAV LNDVLIGFNL N AERDVQAE ...String: MNVSIEEFTH FDFQLVPEPS PLDLVITEPL KNHIEVNGVK SGALLPLPFQ TGIGKTYTAL NFLLQQMLEQ VRSELKEENT GKKSKRLLY YVTDSVDNVV SAKADLLKLI EKQTVKGEPR FTLEQQEYLK AQIVHLPNQS EQLLQCSDAV LNDVLIGFNL N AERDVQAE WSAISGLRRH ASNPEVKISL NRQAGYFYRN LIDRLQKKQK GADRVLLSGS LLASVETLLP GEKIRNGSAH VA FLTTSKF LKGFHNTRSR YSPLRDLSGA VLIIDEIDKQ NQVILSELCK QQAQDLIWAI RTLRANFRDH QLESSPRYDK IED LFEPLR ERLEEFGTNW NLAFAFNTEG ANLNERPVRL FSDRSFTHVS SATHKLSLKS DFLRRKNLIF SDEKVEGSLI EKHG LLTRF VNEADVIYQW FLGTMRKAVF QYWENVRGLE IEVRENRSLE GTFQEAVQSL LTHFNLQEFE SAVYESFDTR GLRQS AGGK ANKLSSSKSY HHTGLKLVEV AHNQGTRDTV NCKASFLNTS PSGVLADMVD AGAVILGISA TARADTVIHN FDFKYL NER LGNKLLSLSR EQKQRVNNYY HSRRNYKDNG VVLTVKYLNS RDAFLDALLE EYKPEARSSH FILNHYLGIA ESEQAFV RS WLSKLLASIK AFISSPDNRY MLSLLNRTLD TTRQNINDFI QFCCDKWAKE FNVKTKTFFG VNADWMRLVG YDEISKHL N TELGKVVVFS TYASMGAGKN PDYAVNLALE GESLISVADV TYSTQLRSDI DSIYLEKPTQ LLLSDDYSHT ANQLCQFHQ ILSLQENGEL SPKSAENWCR QQLMGMSRER SLQQYHQTSD YQSAVRKYIE QAVGRAGRTS LKRKQILLFV DSGLKEILAE ESRDPSLFS HEYVALVNKA KSAGKSIVED RAVRRLFNLA QRNNKDGMLS IKALVHRLHN QPASKSDIQE WQDIRTQLLR Y PTVAFQPE RFNRLYLQSM TKGYYRYQGN LDGDPNSFEF FDRVPYGDMV SEEDCSLATL VQNQYVRPWF ERKGFACSWQ KE ANVMTPI MFTNIYKGAL GEQAVEAVLT AFDFTFEEVP NSIYERFDNR VIFAGIEQPI WLDSKYWKHE GNESSEGYSS KIA LVEEEF GPSKFIYVNA LGDTSKPIRY LNSCFVETSP QLAKVIEIPA LIDDSNADTN RTAVQELIKW LHHS UniProtKB: Helicase/UvrB N-terminal domain-containing protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)