[English] 日本語
 Yorodumi
Yorodumi- EMDB-44232: GluA2 flip Q in complex with TARPgamma2 at pH8, consensus structu... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | GluA2 flip Q in complex with TARPgamma2 at pH8, consensus structure of LBD-TMD-TARPgamma2 | |||||||||||||||
 Map data Map data | GluA2-flip-Q in complex with TARPgamma2 at pH8 (LBD-TMD-STG) | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | AMPA receptor / ionotropic glutamate receptor / ion channel / auxiliary subunit / TRANSPORT PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationPresynaptic depolarization and calcium channel opening / LGI-ADAM interactions / Trafficking of AMPA receptors / eye blink reflex / positive regulation of protein localization to basolateral plasma membrane / cerebellar mossy fiber / postsynaptic neurotransmitter receptor diffusion trapping / regulation of AMPA receptor activity / channel regulator activity / membrane hyperpolarization ...Presynaptic depolarization and calcium channel opening / LGI-ADAM interactions / Trafficking of AMPA receptors / eye blink reflex / positive regulation of protein localization to basolateral plasma membrane / cerebellar mossy fiber / postsynaptic neurotransmitter receptor diffusion trapping / regulation of AMPA receptor activity / channel regulator activity / membrane hyperpolarization / nervous system process / protein targeting to membrane / voltage-gated calcium channel complex / spine synapse / dendritic spine cytoplasm / dendritic spine neck / cellular response to amine stimulus / dendritic spine head / neurotransmitter receptor localization to postsynaptic specialization membrane / Activation of AMPA receptors / ligand-gated monoatomic cation channel activity / perisynaptic space / neuromuscular junction development / Trafficking of GluR2-containing AMPA receptors / AMPA glutamate receptor activity / response to lithium ion / AMPA glutamate receptor clustering / transmission of nerve impulse / kainate selective glutamate receptor activity / immunoglobulin binding / asymmetric synapse / AMPA glutamate receptor complex / regulation of receptor recycling / extracellularly glutamate-gated ion channel activity / cellular response to glycine / ionotropic glutamate receptor complex / membrane depolarization / Unblocking of NMDA receptors, glutamate binding and activation / conditioned place preference / glutamate receptor binding / positive regulation of synaptic transmission / regulation of postsynaptic membrane neurotransmitter receptor levels / voltage-gated calcium channel activity / response to fungicide / regulation of synaptic transmission, glutamatergic / positive regulation of synaptic transmission, glutamatergic / extracellular ligand-gated monoatomic ion channel activity / glutamate-gated receptor activity / cytoskeletal protein binding / cellular response to brain-derived neurotrophic factor stimulus / regulation of long-term synaptic depression / somatodendritic compartment / glutamate-gated calcium ion channel activity / presynaptic active zone membrane / excitatory synapse / ionotropic glutamate receptor signaling pathway / ionotropic glutamate receptor binding / dendrite membrane / ligand-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential / dendrite cytoplasm / positive regulation of excitatory postsynaptic potential / hippocampal mossy fiber to CA3 synapse / dendritic shaft / SNARE binding / synaptic membrane / PDZ domain binding / synaptic transmission, glutamatergic / establishment of protein localization / protein tetramerization / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / regulation of membrane potential / cerebral cortex development / receptor internalization / response to calcium ion / postsynaptic density membrane / modulation of chemical synaptic transmission / Schaffer collateral - CA1 synapse / terminal bouton / synaptic vesicle / long-term synaptic potentiation / synaptic vesicle membrane / amyloid-beta binding / growth cone / presynapse / signaling receptor activity / presynaptic membrane / scaffold protein binding / dendritic spine / chemical synaptic transmission / perikaryon / postsynaptic membrane / neuron projection / postsynaptic density / external side of plasma membrane / axon / neuronal cell body / dendrite / synapse / protein kinase binding / protein-containing complex binding Similarity search - Function | |||||||||||||||
| Biological species |   | |||||||||||||||
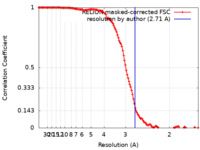
| Method | single particle reconstruction / cryo EM / Resolution: 2.71 Å | |||||||||||||||
 Authors Authors | Nakagawa T / Greger IH | |||||||||||||||
| Funding support |  United States, United States,  United Kingdom, 4 items United Kingdom, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Proton-triggered rearrangement of the AMPA receptor N-terminal domains impacts receptor kinetics and synaptic localization. Authors: Josip Ivica / Nejc Kejzar / Hinze Ho / Imogen Stockwell / Viktor Kuchtiak / Alexander M Scrutton / Terunaga Nakagawa / Ingo H Greger /    Abstract: AMPA glutamate receptors (AMPARs) are ion channel tetramers that mediate the majority of fast excitatory synaptic transmission. They are composed of four subunits (GluA1-GluA4); the GluA2 subunit ...AMPA glutamate receptors (AMPARs) are ion channel tetramers that mediate the majority of fast excitatory synaptic transmission. They are composed of four subunits (GluA1-GluA4); the GluA2 subunit dominates AMPAR function throughout the forebrain. Its extracellular N-terminal domain (NTD) determines receptor localization at the synapse, ensuring reliable synaptic transmission and plasticity. This synaptic anchoring function requires a compact NTD tier, stabilized by a GluA2-specific NTD interface. Here we show that low pH conditions, which accompany synaptic activity, rupture this interface. All-atom molecular dynamics simulations reveal that protonation of an interfacial histidine residue (H208) centrally contributes to NTD rearrangement. Moreover, in stark contrast to their canonical compact arrangement at neutral pH, GluA2 cryo-electron microscopy structures exhibit a wide spectrum of NTD conformations under acidic conditions. We show that the consequences of this pH-dependent conformational control are twofold: rupture of the NTD tier slows recovery from desensitized states and increases receptor mobility at mouse hippocampal synapses. Therefore, a proton-triggered NTD switch will shape both AMPAR location and kinetics, thereby impacting synaptic signal transmission. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_44232.map.gz emd_44232.map.gz | 167 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-44232-v30.xml emd-44232-v30.xml emd-44232.xml emd-44232.xml | 24.9 KB 24.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_44232_fsc.xml emd_44232_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_44232.png emd_44232.png | 241.7 KB | ||
| Filedesc metadata |  emd-44232.cif.gz emd-44232.cif.gz | 7.2 KB | ||
| Others |  emd_44232_additional_1.map.gz emd_44232_additional_1.map.gz emd_44232_additional_2.map.gz emd_44232_additional_2.map.gz emd_44232_half_map_1.map.gz emd_44232_half_map_1.map.gz emd_44232_half_map_2.map.gz emd_44232_half_map_2.map.gz | 166.7 MB 166.4 MB 140.8 MB 140.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-44232 http://ftp.pdbj.org/pub/emdb/structures/EMD-44232 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44232 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44232 | HTTPS FTP |
-Related structure data
| Related structure data |  9b5zMC  9b60C  9b61C  9b63C  9b64C  9b67C  9b68C  9b69C  9b6aC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_44232.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_44232.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | GluA2-flip-Q in complex with TARPgamma2 at pH8 (LBD-TMD-STG) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.82 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: GluA2-flip-Q in complex with TARPgamma2 at pH8 (LBD-TMD-STG,...
| File | emd_44232_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
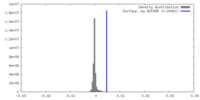
| Annotation | GluA2-flip-Q in complex with TARPgamma2 at pH8 (LBD-TMD-STG, map sharpenning B-factor = -77.8) | ||||||||||||
| Projections & Slices |
| ||||||||||||
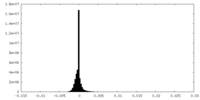
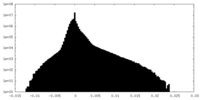
| Density Histograms |
-Additional map: This map is the outcome of the focused...
| File | emd_44232_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
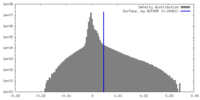
| Annotation | This map is the outcome of the focused refinement of the NTD of the GluA2-flip-Q in complex with TARPgamma2 at pH8 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half1 map: GluA2-flip-Q in complex with TARPgamma2 at...
| File | emd_44232_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
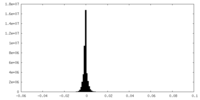
| Annotation | Half1 map: GluA2-flip-Q in complex with TARPgamma2 at pH8 (LBD-TMD-STG) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half2 map: GluA2-flip-Q in complex with TARPgamma2 at...
| File | emd_44232_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
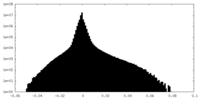
| Annotation | Half2 map: GluA2-flip-Q in complex with TARPgamma2 at pH8 (LBD-TMD-STG) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : GluA2 (flip-Q isoform) in complex with TARPgamma2 at 4:4 stoichiometry
| Entire | Name: GluA2 (flip-Q isoform) in complex with TARPgamma2 at 4:4 stoichiometry |
|---|---|
| Components |
|
-Supramolecule #1: GluA2 (flip-Q isoform) in complex with TARPgamma2 at 4:4 stoichiometry
| Supramolecule | Name: GluA2 (flip-Q isoform) in complex with TARPgamma2 at 4:4 stoichiometry type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Molecular weight | Theoretical: 500 KDa |
-Macromolecule #1: Voltage-dependent calcium channel gamma-2 subunit
| Macromolecule | Name: Voltage-dependent calcium channel gamma-2 subunit / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 35.938746 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGLFDRGVQM LLTTVGAFAA FSLMTIAVGT DYWLYSRGVC KTKSVSENET SKKNEEVMTH SGLWRTCCLE GNFKGLCKQI DHFPEDADY EADTAEYFLR AVRASSIFPI LSVILLFMGG LCIAASEFYK TRHNIILSAG IFFVSAGLSN IIGIIVYISA N AGDPSKSD ...String: MGLFDRGVQM LLTTVGAFAA FSLMTIAVGT DYWLYSRGVC KTKSVSENET SKKNEEVMTH SGLWRTCCLE GNFKGLCKQI DHFPEDADY EADTAEYFLR AVRASSIFPI LSVILLFMGG LCIAASEFYK TRHNIILSAG IFFVSAGLSN IIGIIVYISA N AGDPSKSD SKKNSYSYGW SFYFGALSFI IAEMVGVLAV HMFIDRHKQL RATARATDYL QASAITRIPS YRYRYQRRSR SS SRSTEPS HSRDASPVGV KGFNTLPSTE ISMYTLSRDP LKAATTPTAT YNSDRDNSFL QVHNCIQKDS KDSLHANTAN RRT TPV UniProtKB: Voltage-dependent calcium channel gamma-2 subunit |
-Macromolecule #2: Isoform Flip of Glutamate receptor 2
| Macromolecule | Name: Isoform Flip of Glutamate receptor 2 / type: protein_or_peptide / ID: 2 Details: rat GluA2 flip Q isoform. FLAG epitope tag is located near the C-terminus Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 99.617492 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MQKIMHISVL LSPVLWGLIF GVSSNSIQIG GLFPRGADQE YSAFRVGMVQ FSTSEFRLTP HIDNLEVANS FAVTNAFCSQ FSRGVYAIF GFYDKKSVNT ITSFCGTLHV SFITPSFPTD GTHPFVIQMR PDLKGALLSL IEYYQWDKFA YLYDSDRGLS T LQAVLDSA ...String: MQKIMHISVL LSPVLWGLIF GVSSNSIQIG GLFPRGADQE YSAFRVGMVQ FSTSEFRLTP HIDNLEVANS FAVTNAFCSQ FSRGVYAIF GFYDKKSVNT ITSFCGTLHV SFITPSFPTD GTHPFVIQMR PDLKGALLSL IEYYQWDKFA YLYDSDRGLS T LQAVLDSA AEKKWQVTAI NVGNINNDKK DETYRSLFQD LELKKERRVI LDCERDKVND IVDQVITIGK HVKGYHYIIA NL GFTDGDL LKIQFGGANV SGFQIVDYDD SLVSKFIERW STLEEKEYPG AHTATIKYTS ALTYDAVQVM TEAFRNLRKQ RIE ISRRGN AGDCLANPAV PWGQGVEIER ALKQVQVEGL SGNIKFDQNG KRINYTINIM ELKTNGPRKI GYWSEVDKMV VTLT ELPSG NDTSGLENKT VVVTTILESP YVMMKKNHEM LEGNERYEGY CVDLAAEIAK HCGFKYKLTI VGDGKYGARD ADTKI WNGM VGELVYGKAD IAIAPLTITL VREEVIDFSK PFMSLGISIM IKKPQKSKPG VFSFLDPLAY EIWMCIVFAY IGVSVV LFL VSRFSPYEWH TEEFEDGRET QSSESTNEFG IFNSLWFSLG AFMQQGCDIS PRSLSGRIVG GVWWFFTLII ISSYTAN LA AFLTVERMVS PIESAEDLSK QTEIAYGTLD SGSTKEFFRR SKIAVFDKMW TYMRSAEPSV FVRTTAEGVA RVRKSKGK Y AYLLESTMNE YIEQRKPCDT MKVGGNLDSK GYDIATPKGS SLGTPVNLAV LKLSEQGVLD KLKNKWWYDK GECGAKDSG SKEKTSALSL SNVAGVFYIL VGGLGLAMLV ALIEFCYKSR AEAKRMKVAK NPQNINPSSS QNSQNFATDY KDDDDKEGYN VYGIESVKI UniProtKB: Glutamate receptor 2 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 10 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
Details: Tris adjusted to pH 8 using HCl | ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 21898 / Average electron dose: 52.8 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)