[English] 日本語
 Yorodumi
Yorodumi- EMDB-44010: INF2 at the Barbed End of F-Actin, Leading WH2 focus refinement -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | INF2 at the Barbed End of F-Actin, Leading WH2 focus refinement | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Actin / Filament / Elongation / Ends / CYTOSOLIC PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcytoskeletal motor activator activity / myosin heavy chain binding / tropomyosin binding / regulation of mitochondrial fission / troponin I binding / filamentous actin / mesenchyme migration / actin filament bundle / actin filament bundle assembly / skeletal muscle myofibril ...cytoskeletal motor activator activity / myosin heavy chain binding / tropomyosin binding / regulation of mitochondrial fission / troponin I binding / filamentous actin / mesenchyme migration / actin filament bundle / actin filament bundle assembly / skeletal muscle myofibril / striated muscle thin filament / skeletal muscle thin filament assembly / actin monomer binding / stress fiber / skeletal muscle fiber development / titin binding / actin filament polymerization / filopodium / actin filament / small GTPase binding / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / calcium-dependent protein binding / lamellipodium / actin binding / cell body / hydrolase activity / protein domain specific binding / calcium ion binding / positive regulation of gene expression / perinuclear region of cytoplasm / magnesium ion binding / ATP binding / identical protein binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
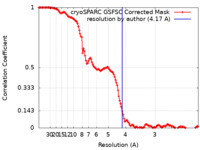
| Method | single particle reconstruction / cryo EM / Resolution: 4.17 Å | |||||||||
 Authors Authors | Palmer NJ / Barrie KR / Dominguez R | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2024 Journal: Nature / Year: 2024Title: Mechanisms of actin filament severing and elongation by formins. Authors: Nicholas J Palmer / Kyle R Barrie / Roberto Dominguez /  Abstract: Humans express 15 formins that play crucial roles in actin-based processes, including cytokinesis, cell motility and mechanotransduction. However, the lack of structures bound to the actin filament ...Humans express 15 formins that play crucial roles in actin-based processes, including cytokinesis, cell motility and mechanotransduction. However, the lack of structures bound to the actin filament (F-actin) has been a major impediment to understanding formin function. Whereas formins are known for their ability to nucleate and elongate F-actin, some formins can additionally depolymerize, sever or bundle F-actin. Two mammalian formins, inverted formin 2 (INF2) and diaphanous 1 (DIA1, encoded by DIAPH1), exemplify this diversity. INF2 shows potent severing activity but elongates weakly whereas DIA1 has potent elongation activity but does not sever. Using cryo-electron microscopy (cryo-EM) we show five structural states of INF2 and two of DIA1 bound to the middle and barbed end of F-actin. INF2 and DIA1 bind differently to these sites, consistent with their distinct activities. The formin-homology 2 and Wiskott-Aldrich syndrome protein-homology 2 (FH2 and WH2, respectively) domains of INF2 are positioned to sever F-actin, whereas DIA1 appears unsuited for severing. These structures also show how profilin-actin is delivered to the fast-growing barbed end, and how this is followed by a transition of the incoming monomer into the F-actin conformation and the release of profilin. Combined, the seven structures presented here provide step-by-step visualization of the mechanisms of F-actin severing and elongation by formins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_44010.map.gz emd_44010.map.gz | 107.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-44010-v30.xml emd-44010-v30.xml emd-44010.xml emd-44010.xml | 19.1 KB 19.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_44010_fsc.xml emd_44010_fsc.xml | 12.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_44010.png emd_44010.png | 31.4 KB | ||
| Masks |  emd_44010_msk_1.map emd_44010_msk_1.map | 216 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-44010.cif.gz emd-44010.cif.gz | 6.2 KB | ||
| Others |  emd_44010_half_map_1.map.gz emd_44010_half_map_1.map.gz emd_44010_half_map_2.map.gz emd_44010_half_map_2.map.gz | 200.2 MB 200.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-44010 http://ftp.pdbj.org/pub/emdb/structures/EMD-44010 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44010 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44010 | HTTPS FTP |
-Related structure data
| Related structure data |  9az4C  9azpC  9azqC  9b03C  9b0kC  9b27C  9b3dC C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_44010.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_44010.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
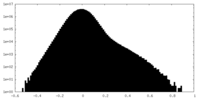
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_44010_msk_1.map emd_44010_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
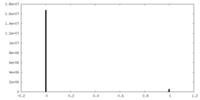
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_44010_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
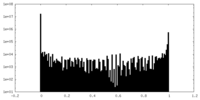
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_44010_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
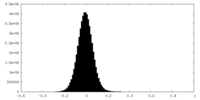
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Actin
| Entire | Name: Actin |
|---|---|
| Components |
|
-Supramolecule #1: Actin
| Supramolecule | Name: Actin / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 264 KDa |
-Supramolecule #2: INF2 Dimer
| Supramolecule | Name: INF2 Dimer / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Actin Filament
| Supramolecule | Name: Actin Filament / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #1 |
|---|
-Macromolecule #1: Actin, alpha skeletal muscle
| Macromolecule | Name: Actin, alpha skeletal muscle / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MCDEDETTAL VCDNGSGLVK AGFAGDDAPR AVFPSIVGRP RHQGVMVGMG QKDSYVGDEA QSKRGILTLK YPIEHGIITN WDDMEKIWHH TFYNELRVAP EEHPTLLTEA PLNPKANREK MTQIMFETFN VPAMYVAIQA VLSLYASGRT TGIVLDSGDG VTHNVPIYEG ...String: MCDEDETTAL VCDNGSGLVK AGFAGDDAPR AVFPSIVGRP RHQGVMVGMG QKDSYVGDEA QSKRGILTLK YPIEHGIITN WDDMEKIWHH TFYNELRVAP EEHPTLLTEA PLNPKANREK MTQIMFETFN VPAMYVAIQA VLSLYASGRT TGIVLDSGDG VTHNVPIYEG YALPHAIMRL DLAGRDLTDY LMKILTERGY SFVTTAEREI VRDIKEKLCY VALDFENEMA TAASSSSLEK SYELPDGQVI TIGNERFRCP ETLFQPSFIG MESAGIHETT YNSIMKCDID IRKDLYANNV MSGGTTMYPG IADRMQKEIT ALAPSTMKIK IIAPPERKYS VWIGGSILAS LSTFQQMWIT KQEYDEAGPS IVHRKCF UniProtKB: Actin, alpha skeletal muscle |
-Macromolecule #2: INF2
| Macromolecule | Name: INF2 / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: MSVKEGAQRK WAALKEKLGP QDSDPTEANL ESADPELCIR LLQMPSVVNY SGLRKRLEGS DGGWMVQFLE QSGLDLLLEA LARLSGRGVA RISDALLQLT CVSCVRAVMN SRQGIEYILS NQGYVRQLSQ ALDTSNVMVK KQVFELLAAL CIYSPEGHVL TLDALDHYKT ...String: MSVKEGAQRK WAALKEKLGP QDSDPTEANL ESADPELCIR LLQMPSVVNY SGLRKRLEGS DGGWMVQFLE QSGLDLLLEA LARLSGRGVA RISDALLQLT CVSCVRAVMN SRQGIEYILS NQGYVRQLSQ ALDTSNVMVK KQVFELLAAL CIYSPEGHVL TLDALDHYKT VCSQQYRFSI VMNELSGSDN VPYVVTLLSV INAVILGPED LRARTQLRNE FIGLQLLDVL ARLRDLEDAD LLIQLEAFEE AKAEDEEELL RVSGGVDMSS HQEVFASLFH KVSCSPVSAQ LLSVLQGLLH LEPTLRSSQL LWEALESLVN RAVLLASDAQ ECTLEEVVER LLSVKGRPRP SPLVKAHKSV QANLDQSQRG SSPQNTTTPK PSVEGQQPAA AAACEPVDHA QSESILKVSQ PRALEQQAST PPPPPPPPLL PGSSAEPPPP PPPPPLPSVG AKALPTAPPP PPLPGLGAMA PPAPPLPPPL PGSCEFLPPP PPPLPGLGCP PPPPPLLPGM GWGPPPPPPP LLPCTCSPPV AGGMEEVIVA QVDHGLGSAW VPSHRRVNPP TLRMKKLNWQ KLPSNVAREH NSMWASLSSP DAEAVEPDFS SIERLFSFPA AKPKEPTMVA PRARKEPKEI TFLDAKKSLN LNIFLKQFKC SNEEVAAMIR AGDTTKFDVE VLKQLLKLLP EKHEIENLRA FTEERAKLAS ADHFYLLLLA IPCYQLRIEC MLLCEGAAAV LDMVRPKAQL VLAACESLLT SRQLPIFCQL ILRIGNFLNY GSHTGDADGF KISTLLKLTE TKSQQNRVTL LHHVLEEAEK SHPDLLQLPR DLEQPSQAAG INLEIIRSEA SSNLKKLLET ERKVSASVAE VQEQYTERLQ ASISAFRALD ELFEAIEQKQ RELADYLCED AQQLSLEDTF STMKAFRDLF LRALKENKDR KEQAAKAERR KQQLAEEEAR RPRGEDGKPV RKGPGKQEEV CVIDALLADI RKGFQLRKTA RGRGDTDGGS KAASMDPPRA TEPVATSNPA GDPVGSTRCP ASEPGLDATT ASESRGWDLV DAVTPGPQPT LEQLEEGGPR PLERRSSWYV DASDVLTTED PQCPQPLEGA WPVTLGDAQA LKPLKFSSNQ PPAAGSSRQD AKDPTSLLGV LQAEADSTSE GLEDAVHSRG ARPPAAGPGG DEDEDEEDTA PESALDTSLD KSFSEDAVTD SSGSGTLPRA RGRASKGTGK RRKKRPSRSQ EEVPPDSDDN KTKKLCVIQ UniProtKB: Inverted formin-2 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.45 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 100 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 120 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 41926 / Average electron dose: 44.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)