+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4222 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | NCP interactions : Class A2 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species | ||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 9.0 Å | |||||||||
 Authors Authors | Halic M / Bilokapic S | |||||||||
 Citation Citation |  Journal: Sci Rep / Year: 2018 Journal: Sci Rep / Year: 2018Title: Cryo-EM of nucleosome core particle interactions in trans. Authors: Silvija Bilokapic / Mike Strauss / Mario Halic /  Abstract: Nucleosomes, the basic unit of chromatin, are repetitively spaced along DNA and regulate genome expression and maintenance. The long linear chromatin molecule is extensively condensed to fit DNA ...Nucleosomes, the basic unit of chromatin, are repetitively spaced along DNA and regulate genome expression and maintenance. The long linear chromatin molecule is extensively condensed to fit DNA inside the nucleus. How distant nucleosomes interact to build tertiary chromatin structure remains elusive. In this study, we used cryo-EM to structurally characterize different states of long range nucleosome core particle (NCP) interactions. Our structures show that NCP pairs can adopt multiple conformations, but, commonly, two NCPs are oriented with the histone octamers facing each other. In this conformation, the dyad of both nucleosome core particles is facing the same direction, however, the NCPs are laterally shifted and tilted. The histone octamer surface and histone tails in trans NCP pairs remain accessible to regulatory proteins. The overall conformational flexibility of the NCP pair suggests that chromatin tertiary structure is dynamic and allows access of various chromatin modifying machineries to nucleosomes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4222.map.gz emd_4222.map.gz | 25.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4222-v30.xml emd-4222-v30.xml emd-4222.xml emd-4222.xml | 7.5 KB 7.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_4222.png emd_4222.png | 84.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4222 http://ftp.pdbj.org/pub/emdb/structures/EMD-4222 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4222 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4222 | HTTPS FTP |
-Validation report
| Summary document |  emd_4222_validation.pdf.gz emd_4222_validation.pdf.gz | 201.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_4222_full_validation.pdf.gz emd_4222_full_validation.pdf.gz | 200.4 KB | Display | |
| Data in XML |  emd_4222_validation.xml.gz emd_4222_validation.xml.gz | 6.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4222 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4222 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4222 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4222 | HTTPS FTP |
-Related structure data
| Related structure data |  4221C  4223C  4224C  4226C  4227C  4228C  4229C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4222.map.gz / Format: CCP4 / Size: 42.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4222.map.gz / Format: CCP4 / Size: 42.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.4 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
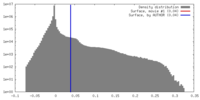
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Nucleosome
| Entire | Name: Nucleosome |
|---|---|
| Components |
|
-Supramolecule #1: Nucleosome
| Supramolecule | Name: Nucleosome / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism: |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 200 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Average electron dose: 100.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 9.0 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 1.4) / Number images used: 5000 |
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)