[English] 日本語
 Yorodumi
Yorodumi- EMDB-41757: Structure of full length LRRK2 bound to GZD-824 (I2020T mutant) -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of full length LRRK2 bound to GZD-824 (I2020T mutant) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Kinase inhibitors / Kinase / GTPases / PROTEIN BINDING | |||||||||
| Function / homology |  Function and homology information Function and homology informationcaveola neck / : / beta-catenin destruction complex binding / regulation of branching morphogenesis of a nerve / Wnt signalosome assembly / negative regulation of motile cilium assembly / regulation of kidney size / regulation of cell projection organization / tangential migration from the subventricular zone to the olfactory bulb / GTP-dependent protein kinase activity ...caveola neck / : / beta-catenin destruction complex binding / regulation of branching morphogenesis of a nerve / Wnt signalosome assembly / negative regulation of motile cilium assembly / regulation of kidney size / regulation of cell projection organization / tangential migration from the subventricular zone to the olfactory bulb / GTP-dependent protein kinase activity / regulation of SNARE complex assembly / regulation of neuroblast proliferation / regulation of ER to Golgi vesicle-mediated transport / protein localization to endoplasmic reticulum exit site / peroxidase inhibitor activity / negative regulation of late endosome to lysosome transport / regulation of mitochondrial depolarization / : / positive regulation of dopamine receptor signaling pathway / amphisome / regulation of synaptic vesicle transport / : / regulation of CAMKK-AMPK signaling cascade / co-receptor binding / negative regulation of GTPase activity / regulation of dopamine receptor signaling pathway / positive regulation of microglial cell activation / regulation of retrograde transport, endosome to Golgi / regulation of neuron maturation / cytoplasmic side of mitochondrial outer membrane / cellular response to curcumin / negative regulation of autophagosome assembly / olfactory bulb development / positive regulation of synaptic vesicle endocytosis / JUN kinase kinase kinase activity / multivesicular body, internal vesicle / negative regulation of excitatory postsynaptic potential / regulation of cAMP/PKA signal transduction / striatum development / neuron projection arborization / mitochondrion localization / regulation of dendritic spine morphogenesis / protein localization to mitochondrion / cellular response to dopamine / positive regulation of mitochondrial outer membrane permeabilization involved in apoptotic signaling pathway / endoplasmic reticulum organization / positive regulation of protein autoubiquitination / negative regulation of protein processing / Wnt signalosome / positive regulation of programmed cell death / GTP metabolic process / regulation of canonical Wnt signaling pathway / syntaxin-1 binding / regulation of reactive oxygen species metabolic process / Golgi-associated vesicle / PTK6 promotes HIF1A stabilization / lysosome organization / clathrin binding / negative regulation of macroautophagy / regulation of mitochondrial fission / regulation of synaptic vesicle exocytosis / Lewy body / regulation of locomotion / intracellular distribution of mitochondria / Golgi organization / neuromuscular junction development / protein kinase A binding / microvillus / autolysosome / exploration behavior / locomotory exploration behavior / endoplasmic reticulum exit site / negative regulation of Notch signaling pathway / MAP kinase kinase kinase activity / regulation of synaptic vesicle endocytosis / canonical Wnt signaling pathway / negative regulation of endoplasmic reticulum stress-induced intrinsic apoptotic signaling pathway / Rho protein signal transduction / regulation of synaptic transmission, glutamatergic / presynaptic cytosol / neuron projection morphogenesis / phagocytic vesicle / JNK cascade / cellular response to manganese ion / positive regulation of autophagy / dendrite cytoplasm / positive regulation of protein ubiquitination / cellular response to starvation / GTPase activator activity / regulation of autophagy / determination of adult lifespan / SNARE binding / cellular response to reactive oxygen species / mitochondrion organization / trans-Golgi network / calcium-mediated signaling / excitatory postsynaptic potential / regulation of protein stability / tubulin binding / regulation of membrane potential Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
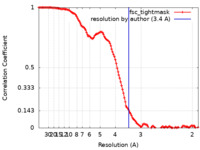
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Villagran-Suarez A / Sanz-Murillo M / Alegrio-Louro J / Leschziner A | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2023 Journal: Sci Adv / Year: 2023Title: Inhibition of Parkinson's disease-related LRRK2 by type I and type II kinase inhibitors: Activity and structures. Authors: Marta Sanz Murillo / Amalia Villagran Suarez / Verena Dederer / Deep Chatterjee / Jaime Alegrio Louro / Stefan Knapp / Sebastian Mathea / Andres E Leschziner /   Abstract: Mutations in leucine-rich repeat kinase 2 (LRRK2) are a common cause of familial Parkinson's disease (PD) and a risk factor for the sporadic form. Increased kinase activity was shown in patients with ...Mutations in leucine-rich repeat kinase 2 (LRRK2) are a common cause of familial Parkinson's disease (PD) and a risk factor for the sporadic form. Increased kinase activity was shown in patients with both familial and sporadic PD, making LRRK2 kinase inhibitors a major focus of drug development efforts. Although much progress has been made in understanding the structural biology of LRRK2, there are no available structures of LRRK2 inhibitor complexes. To this end, we solved cryo-electron microscopy structures of LRRK2, wild-type and PD-linked mutants, bound to the LRRK2-specific type I inhibitor MLi-2 and the broad-spectrum type II inhibitor GZD-824. Our structures revealed an active-like LRRK2 kinase in the type I inhibitor complex, and an inactive DYG-out in the type II inhibitor complex. Our structural analysis also showed how inhibitor-induced conformational changes in LRRK2 are affected by its autoinhibitory N-terminal repeats. The structures provide a template for the rational development of LRRK2 kinase inhibitors covering both canonical inhibitor binding modes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41757.map.gz emd_41757.map.gz | 168.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41757-v30.xml emd-41757-v30.xml emd-41757.xml emd-41757.xml | 19.1 KB 19.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_41757_fsc.xml emd_41757_fsc.xml | 13.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_41757.png emd_41757.png | 45.3 KB | ||
| Filedesc metadata |  emd-41757.cif.gz emd-41757.cif.gz | 7.7 KB | ||
| Others |  emd_41757_half_map_1.map.gz emd_41757_half_map_1.map.gz emd_41757_half_map_2.map.gz emd_41757_half_map_2.map.gz | 165.1 MB 165.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41757 http://ftp.pdbj.org/pub/emdb/structures/EMD-41757 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41757 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41757 | HTTPS FTP |
-Related structure data
| Related structure data |  8tzfMC  8txzC  8tyqC  8tzbC  8tzcC  8tzeC  8tzgC  8tzhC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_41757.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41757.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.95 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_41757_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_41757_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of LRRK2 with GZD-824 and Darpin bound
| Entire | Name: Complex of LRRK2 with GZD-824 and Darpin bound |
|---|---|
| Components |
|
-Supramolecule #1: Complex of LRRK2 with GZD-824 and Darpin bound
| Supramolecule | Name: Complex of LRRK2 with GZD-824 and Darpin bound / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Leucine-rich repeat serine/threonine-protein kinase 2
| Macromolecule | Name: Leucine-rich repeat serine/threonine-protein kinase 2 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 286.450719 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASGSCQGCE EDEETLKKLI VRLNNVQEGK QIETLVQILE DLLVFTYSER ASKLFQGKNI HVPLLIVLDS YMRVASVQQV GWSLLCKLI EVCPGTMQSL MGPQDVGNDW EVLGVHQLIL KMLTVHNASV NLSVIGLKTL DLLLTSGKIT LLILDEESDI F MLIFDAMH ...String: MASGSCQGCE EDEETLKKLI VRLNNVQEGK QIETLVQILE DLLVFTYSER ASKLFQGKNI HVPLLIVLDS YMRVASVQQV GWSLLCKLI EVCPGTMQSL MGPQDVGNDW EVLGVHQLIL KMLTVHNASV NLSVIGLKTL DLLLTSGKIT LLILDEESDI F MLIFDAMH SFPANDEVQK LGCKALHVLF ERVSEEQLTE FVENKDYMIL LSALTNFKDE EEIVLHVLHC LHSLAIPCNN VE VLMSGNV RCYNIVVEAM KAFPMSERIQ EVSCCLLHRL TLGNFFNILV LNEVHEFVVK AVQQYPENAA LQISALSCLA LLT ETIFLN QDLEEKNENQ ENDDEGEEDK LFWLEACYKA LTWHRKNKHV QEAACWALNN LLMYQNSLHE KIGDEDGHFP AHRE VMLSM LMHSSSKEVF QASANALSTL LEQNVNFRKI LLSKGIHLNV LELMQKHIHS PEVAESGCKM LNHLFEGSNT SLDIM AAVV PKILTVMKRH ETSLPVQLEA LRAILHFIVP GMPEESREDT EFHHKLNMVK KQCFKNDIHK LVLAALNRFI GNPGIQ KCG LKVISSIVHF PDALEMLSLE GAMDSVLHTL QMYPDDQEIQ CLGLSLIGYL ITKKNVFIGT GHLLAKILVS SLYRFKD VA EIQTKGFQTI LAILKLSASF SKLLVHHSFD LVIFHQMSSN IMEQKDQQFL NLCCKCFAKV AMDDYLKNVM LERACDQN N SIMVECLLLL GADANQAKEG SSLICQVCEK ESSPKLVELL LNSGSREQDV RKALTISIGK GDSQIISLLL RRLALDVAN NSICLGGFCI GKVEPSWLGP LFPDKTSNLR KQTNIASTLA RMVIRYQMKS AVEEGTASGS DGNFSEDVLS KFDEWTFIPD SSMDSVFAQ SDDLDSEGSE GSFLVKKKSN SISVGEFYRD AVLQRCSPNL QRHSNSLGPI FDHEDLLKRK RKILSSDDSL R SSKLQSHM RHSDSISSLA SEREYITSLD LSANELRDID ALSQKCCISV HLEHLEKLEL HQNALTSFPQ QLCETLKSLT HL DLHSNKF TSFPSYLLKM SCIANLDVSR NDIGPSVVLD PTVKCPTLKQ FNLSYNQLSF VPENLTDVVE KLEQLILEGN KIS GICSPL RLKELKILNL SKNHISSLSE NFLEACPKVE SFSARMNFLA AMPFLPPSMT ILKLSQNKFS CIPEAILNLP HLRS LDMSS NDIQYLPGPA HWKSLNLREL LFSHNQISIL DLSEKAYLWS RVEKLHLSHN KLKEIPPEIG CLENLTSLDV SYNLE LRSF PNEMGKLSKI WDLPLDELHL NFDFKHIGCK AKDIIRFLQQ RLKKAVPYNR MKLMIVGNTG SGKTTLLQQL MKTKKS DLG MQSATVGIDV KDWPIQIRDK RKRDLVLNVW DFAGREEFYS THPHFMTQRA LYLAVYDLSK GQAEVDAMKP WLFNIKA RA SSSPVILVGT HLDVSDEKQR KACMSKITKE LLNKRGFPAI RDYHFVNATE ESDALAKLRK TIINESLNFK IRDQLVVG Q LIPDCYVELE KIILSERKNV PIEFPVIDRK RLLQLVRENQ LQLDENELPH AVHFLNESGV LLHFQDPALQ LSDLYFVEP KWLCKIMAQI LTVKVEGCPK HPKGIISRRD VEKFLSKKRK FPKNYMSQYF KLLEKFQIAL PIGEEYLLVP SSLSDHRPVI ELPHCENSE IIIRLYEMPY FPMGFWSRLI NRLLEISPYM LSGRERALRP NRMYWRQGIY LNWSPEAYCL VGSEVLDNHP E SFLKITVP SCRKGCILLG QVVDHIDSLM EEWFPGLLEI DICGEGETLL KKWALYSFND GEEHQKILLD DLMKKAEEGD LL VNPDQPR LTIPISQIAP DLILADLPRN IMLNNDELEF EQAPEFLLGD GSFGSVYRAA YEGEEVAVKI FNKHTSLRLL RQE LVVLCH LHHPSLISLL AAGIRPRMLV MELASKGSLD RLLQQDKASL TRTLQHRIAL HVADGLRYLH SAMIIYRDLK PHNV LLFTL YPNAAIIAKI ADYGTAQYCC RMGIKTSEGT PGFRAPEVAR GNVIYNQQAD VYSFGLLLYD ILTTGGRIVE GLKFP NEFD ELEIQGKLPD PVKEYGCAPW PMVEKLIKQC LKENPQERPT SAQVFDILNS AELVCLTRRI LLPKNVIVEC MVATHH NSR NASIWLGCGH TDRGQLSFLD LNTEGYTSEE VADSRILCLA LVHLPVEKES WIVSGTQSGT LLVINTEDGK KRHTLEK MT DSVTCLYCNS FSKQSKQKNF LLVGTADGKL AIFEDKTVKL KGAAPLKILN IGNVSTPLMC LSESTNSTER NVMWGGCG T KIFSFSNDFT IQKLIETRTS QLFSYAAFSD SNIITVVVDT ALYIAKQNSP VVEVWDKKTE KLCGLIDCVH FLREVMVKE NKESKHKMSY SGRVKTLCLQ KNTALWIGTG GGHILLLDLS TRRLIRVIYN FCNSVRVMMT AQLGSLKNVM LVLGYNRKNT EGTQKQKEI QSCLTVWDIN LPHEVQNLEK HIEVRKELAE KMRRTSVE UniProtKB: Leucine-rich repeat serine/threonine-protein kinase 2 |
-Macromolecule #2: designed ankyrin repeat proteins E11
| Macromolecule | Name: designed ankyrin repeat proteins E11 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 19.766912 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MRGSHHHHHH HHGSDLGKKL LEAARAGQDD EVRILMANGA DVNATDEAGV TPLHLAADSG HLEIVEVLLK TGADVNAWDH YGFTPLHLA AHVGHLEIVE VLLKAGADVN AQDHAGWTPL HLAALYGHLE IVEVLLKHGA DVNAQDMWGE TPFDLAIDNG N EDIAEVLQ KAAKLNDYKD DDDK |
-Macromolecule #3: GUANOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 1 / Formula: GDP |
|---|---|
| Molecular weight | Theoretical: 443.201 Da |
| Chemical component information |  ChemComp-GDP: |
-Macromolecule #4: 4-methyl-N-{4-[(4-methylpiperazin-1-yl)methyl]-3-(trifluoromethyl...
| Macromolecule | Name: 4-methyl-N-{4-[(4-methylpiperazin-1-yl)methyl]-3-(trifluoromethyl)phenyl}-3-[(1H-pyrazolo[3,4-b]pyridin-5-yl)ethynyl]benzamide type: ligand / ID: 4 / Number of copies: 1 / Formula: T3X |
|---|---|
| Molecular weight | Theoretical: 532.559 Da |
| Chemical component information |  ChemComp-T3X: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.43 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: UltrAuFoil R2/2 / Material: GOLD / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Number grids imaged: 1 / Number real images: 4102 / Average electron dose: 55.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 150000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)