+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | TCR in nanodisc ND-II | |||||||||||||||||||||
 Map data Map data | unsharpened map | |||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | T-cell receptor / TCR / 1G4 / nanodisc / CD3 / IMMUNE SYSTEM | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationdetection of tumor cell / regulation of lymphocyte apoptotic process / gamma-delta T cell receptor complex / Fc-gamma receptor III complex / T cell anergy / positive regulation of cell-cell adhesion mediated by integrin / positive regulation of T cell anergy / Fc-gamma receptor signaling pathway / gamma-delta T cell activation / CD4-positive, alpha-beta T cell proliferation ...detection of tumor cell / regulation of lymphocyte apoptotic process / gamma-delta T cell receptor complex / Fc-gamma receptor III complex / T cell anergy / positive regulation of cell-cell adhesion mediated by integrin / positive regulation of T cell anergy / Fc-gamma receptor signaling pathway / gamma-delta T cell activation / CD4-positive, alpha-beta T cell proliferation / negative thymic T cell selection / positive regulation of CD4-positive, alpha-beta T cell proliferation / alpha-beta T cell receptor complex / positive thymic T cell selection / positive regulation of protein localization to cell surface / signal complex assembly / Nef and signal transduction / T cell mediated cytotoxicity directed against tumor cell target / positive regulation of cell-matrix adhesion / T cell receptor complex / smoothened signaling pathway / Translocation of ZAP-70 to Immunological synapse / Phosphorylation of CD3 and TCR zeta chains / positive regulation of interleukin-4 production / establishment or maintenance of cell polarity / dendrite development / protein complex oligomerization / alpha-beta T cell activation / Generation of second messenger molecules / FCGR activation / immunological synapse / Co-inhibition by PD-1 / Role of phospholipids in phagocytosis / T cell receptor binding / T cell costimulation / positive regulation of T cell proliferation / positive regulation of interleukin-2 production / positive regulation of calcium-mediated signaling / FCGR3A-mediated IL10 synthesis / cell surface receptor protein tyrosine kinase signaling pathway / protein tyrosine kinase binding / T cell activation / bioluminescence / cerebellum development / negative regulation of smoothened signaling pathway / generation of precursor metabolites and energy / FCGR3A-mediated phagocytosis / apoptotic signaling pathway / clathrin-coated endocytic vesicle membrane / calcium-mediated signaling / Regulation of actin dynamics for phagocytic cup formation / SH3 domain binding / positive regulation of type II interferon production / Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell / cell-cell junction / transmembrane signaling receptor activity / Downstream TCR signaling / Cargo recognition for clathrin-mediated endocytosis / protein transport / T cell receptor signaling pathway / signaling receptor activity / signaling receptor complex adaptor activity / Clathrin-mediated endocytosis / cell body / protein-containing complex assembly / regulation of apoptotic process / adaptive immune response / dendritic spine / cell surface receptor signaling pathway / G protein-coupled receptor signaling pathway / protein heterodimerization activity / negative regulation of gene expression / external side of plasma membrane / positive regulation of gene expression / protein kinase binding / endoplasmic reticulum / Golgi apparatus / protein homodimerization activity / identical protein binding / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  human respiratory syncytial virus / human respiratory syncytial virus /  | |||||||||||||||||||||
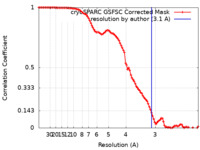
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||||||||||||||
 Authors Authors | Notti RQ / Walz T | |||||||||||||||||||||
| Funding support |  United States, 6 items United States, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: bioRxiv / Year: 2024 Journal: bioRxiv / Year: 2024Title: The resting and ligand-bound states of the membrane-embedded human T-cell receptor-CD3 complex. Authors: Ryan Q Notti / Fei Yi / Søren Heissel / Martin W Bush / Zaki Molvi / Pujita Das / Henrik Molina / Christopher A Klebanoff / Thomas Walz Abstract: The T-cell receptor (TCR) initiates T-lymphocyte activation, but mechanistic questions remain( ). Here, we present cryogenic electron microscopy structures for the unliganded and human leukocyte ...The T-cell receptor (TCR) initiates T-lymphocyte activation, but mechanistic questions remain( ). Here, we present cryogenic electron microscopy structures for the unliganded and human leukocyte antigen (HLA)-bound human TCR-CD3 complex in nanodiscs that provide a native-like lipid environment. Distinct from the "open and extended" conformation seen in detergent( ), the unliganded TCR-CD3 in nanodiscs adopts two related "closed and compacted" conformations that represent its physiologic resting state . By contrast, the HLA-bound complex adopts the open and extended conformation, and conformation-locking disulfide mutants show that ectodomain opening is necessary for maximal ligand-dependent T-cell activation. Together, these results reveal allosteric conformational change during TCR activation and highlight the importance of native-like lipid environments for membrane protein structure determination. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41660.map.gz emd_41660.map.gz | 51.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41660-v30.xml emd-41660-v30.xml emd-41660.xml emd-41660.xml | 27.5 KB 27.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_41660_fsc.xml emd_41660_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_41660.png emd_41660.png | 43.7 KB | ||
| Filedesc metadata |  emd-41660.cif.gz emd-41660.cif.gz | 7.7 KB | ||
| Others |  emd_41660_additional_1.map.gz emd_41660_additional_1.map.gz emd_41660_additional_2.map.gz emd_41660_additional_2.map.gz emd_41660_half_map_1.map.gz emd_41660_half_map_1.map.gz emd_41660_half_map_2.map.gz emd_41660_half_map_2.map.gz | 50.8 MB 111 MB 95.7 MB 115.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41660 http://ftp.pdbj.org/pub/emdb/structures/EMD-41660 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41660 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41660 | HTTPS FTP |
-Validation report
| Summary document |  emd_41660_validation.pdf.gz emd_41660_validation.pdf.gz | 412.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_41660_full_validation.pdf.gz emd_41660_full_validation.pdf.gz | 412 KB | Display | |
| Data in XML |  emd_41660_validation.xml.gz emd_41660_validation.xml.gz | 6.5 KB | Display | |
| Data in CIF |  emd_41660_validation.cif.gz emd_41660_validation.cif.gz | 7.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41660 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41660 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41660 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41660 | HTTPS FTP |
-Related structure data
| Related structure data |  8tw6MC  8tw4C  9bbcC  9c3eC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_41660.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41660.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: B-factor sharpened
| File | emd_41660_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | B-factor sharpened | ||||||||||||
| Projections & Slices |
| ||||||||||||
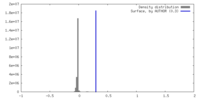
| Density Histograms |
-Additional map: unsharpened map
| File | emd_41660_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
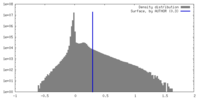
| Density Histograms |
-Half map: unsharpened map
| File | emd_41660_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
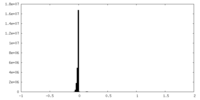
| Density Histograms |
-Half map: unsharpened map
| File | emd_41660_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
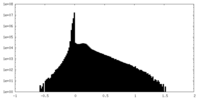
| Density Histograms |
- Sample components
Sample components
-Entire : TCR hetero-octamer in lipid nanodisc
| Entire | Name: TCR hetero-octamer in lipid nanodisc |
|---|---|
| Components |
|
-Supramolecule #1: TCR hetero-octamer in lipid nanodisc
| Supramolecule | Name: TCR hetero-octamer in lipid nanodisc / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: T-cell surface glycoprotein CD3 zeta chain, GFP fusion protein,GFP
| Macromolecule | Name: T-cell surface glycoprotein CD3 zeta chain, GFP fusion protein,GFP type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  human respiratory syncytial virus human respiratory syncytial virus |
| Molecular weight | Theoretical: 46.650879 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MKWKALFTAA ILQAQLPITE AQSFGLLDPK LCYLLDGILF IYGVILTALF LRVKFSRSAD APAYQQGQNQ LYNELNLGRR EEYDVLDKR RGRDPEMGGK PQRRKNPQEG LYNELQKDKM AEAYSEIGMK GERRRGKGHD GLYQGLSTAT KDTYDALHMQ A LPPRASLE ...String: MKWKALFTAA ILQAQLPITE AQSFGLLDPK LCYLLDGILF IYGVILTALF LRVKFSRSAD APAYQQGQNQ LYNELNLGRR EEYDVLDKR RGRDPEMGGK PQRRKNPQEG LYNELQKDKM AEAYSEIGMK GERRRGKGHD GLYQGLSTAT KDTYDALHMQ A LPPRASLE VLFQGPVSKG EELFTGVVPI LVELDGDVNG HKFSVSGEGE GDATYGKLTL KFICTTGKLP VPWPTLVTTL TY GVQCFSR YPDHMKQHDF FKSAMPEGYV QERTIFFKDD GNYKTRAEVK FEGDTLVNRI ELKGIDFKED GNILGHKLEY NYN SHNVYI MADKQKNGIK VNFKIRHNIE DGSVQLADHY QQNTPIGDGP VLLPDNHYLS TQSKLSKDPN EKRDHMVLLE FVTA AGITL GMDELYK UniProtKB: T-cell surface glycoprotein CD3 zeta chain, GFP |
-Macromolecule #2: TCR alpha
| Macromolecule | Name: TCR alpha / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 30.388189 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: METLLGLLIL WLQLQWVSSK QEVTQIPAAL SVPEGENLVL NCSFTDSAIY NLQWFRQDPG KGLTSLLLIQ SSQREQTSGR LNASLDKSS GRSTLYIAAS QPGDSATYLC AVRPLYGGSY IPTFGRGTSL IVHPYIQNPD PAVYQLRDSK SSDKSVCLFT D FDSQTNVS ...String: METLLGLLIL WLQLQWVSSK QEVTQIPAAL SVPEGENLVL NCSFTDSAIY NLQWFRQDPG KGLTSLLLIQ SSQREQTSGR LNASLDKSS GRSTLYIAAS QPGDSATYLC AVRPLYGGSY IPTFGRGTSL IVHPYIQNPD PAVYQLRDSK SSDKSVCLFT D FDSQTNVS QSKDSDVYIT DKTVLDMRSM DFKSNSAVAW SNKSDFACAN AFNNSIIPED TFFPSPESSC DVKLVEKSFE TD TNLNFQN LSVIGFRILL LKVAGFNLLM TLRLWSS |
-Macromolecule #3: T cell receptor beta variable 6-5,T cell receptor beta chain MC.7...
| Macromolecule | Name: T cell receptor beta variable 6-5,T cell receptor beta chain MC.7.G5,MCHERRY type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 62.042883 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSIGLLCCAA LSLLWAGPVN AGVTQTPKFQ VLKTGQSMTL QCAQDMNHEY MSWYRQDPGM GLRLIHYSVG AGITDQGEVP NGYNVSRST TEDFPLRLLS AAPSQTSVYF CASSYVGNTG ELFFGEGSRL TVLEDLKNVF PPEVAVFEPS EAEISHTQKA T LVCLATGF ...String: MSIGLLCCAA LSLLWAGPVN AGVTQTPKFQ VLKTGQSMTL QCAQDMNHEY MSWYRQDPGM GLRLIHYSVG AGITDQGEVP NGYNVSRST TEDFPLRLLS AAPSQTSVYF CASSYVGNTG ELFFGEGSRL TVLEDLKNVF PPEVAVFEPS EAEISHTQKA T LVCLATGF YPDHVELSWW VNGKEVHSGV STDPQPLKEQ PALNDSRYCL SSRLRVSATF WQNPRNHFRC QVQFYGLSEN DE WTQDRAK PVTQIVSAEA WGRADCGFTS ESYQQGVLSA TILYEILLGK ATLYAVLVSA LVLMAMVKRK DSRGASLEVL FQG PVSKGE EDNMAIIKEF MRFKVHMEGS VNGHEFEIEG EGEGRPYEGT QTAKLKVTKG GPLPFAWDIL SPQFMYGSKA YVKH PADIP DYLKLSFPEG FKWERVMNFE DGGVVTVTQD SSLQDGEFIY KVKLRGTNFP SDGPVMQKKT MGWEASSERM YPEDG ALKG EIKQRLKLKD GGHYDAEVKT TYKAKKPVQL PGAYNVNIKL DITSHNEDYT IVEQYERAEG RHSTGGMDEL YK UniProtKB: T cell receptor beta variable 6-5, T cell receptor beta chain MC.7.G5, MCHERRY |
-Macromolecule #4: T-cell surface glycoprotein CD3 epsilon chain
| Macromolecule | Name: T-cell surface glycoprotein CD3 epsilon chain / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.174227 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MQSGTHWRVL GLCLLSVGVW GQDGNEEMGG ITQTPYKVSI SGTTVILTCP QYPGSEILWQ HNDKNIGGDE DDKNIGSDED HLSLKEFSE LEQSGYYVCY PRGSKPEDAN FYLYLRARVC ENCMEMDVMS VATIVIVDIC ITGGLLLLVY YWSKNRKAKA K PVTRGAGA ...String: MQSGTHWRVL GLCLLSVGVW GQDGNEEMGG ITQTPYKVSI SGTTVILTCP QYPGSEILWQ HNDKNIGGDE DDKNIGSDED HLSLKEFSE LEQSGYYVCY PRGSKPEDAN FYLYLRARVC ENCMEMDVMS VATIVIVDIC ITGGLLLLVY YWSKNRKAKA K PVTRGAGA GGRQRGQNKE RPPPVPNPDY EPIRKGQRDL YSGLNQRRI UniProtKB: T-cell surface glycoprotein CD3 epsilon chain |
-Macromolecule #5: T-cell surface glycoprotein CD3 delta chain
| Macromolecule | Name: T-cell surface glycoprotein CD3 delta chain / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 18.949537 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MEHSTFLSGL VLATLLSQVS PFKIPIEELE DRVFVNCNTS ITWVEGTVGT LLSDITRLDL GKRILDPRGI YRCNGTDIYK DKESTVQVH YRMCQSCVEL DPATVAGIIV TDVIATLLLA LGVFCFAGHE TGRLSGAADT QALLRNDQVY QPLRDRDDAQ Y SHLGGNWA RNK UniProtKB: T-cell surface glycoprotein CD3 delta chain |
-Macromolecule #6: T-cell surface glycoprotein CD3 gamma chain
| Macromolecule | Name: T-cell surface glycoprotein CD3 gamma chain / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 21.598633 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MEQGKGLAVL ILAIILLQGT LAQSIKGNHL VKVYDYQEDG SVLLTCDAEA KNITWFKDGK MIGFLTEDKK KWNLGSNAKD PRGMYQCKG SQNKSKPLQV YYRMCQNCIE LNAATISGFL FAEIVSIFVL AVGVYFIAGQ DGVRQSRASD KQTLLPNDQL Y QPLKDRED DQYSHLQGNQ LRRNHHHHHH HH UniProtKB: T-cell surface glycoprotein CD3 gamma chain |
-Macromolecule #8: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 8 / Number of copies: 2 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Macromolecule #9: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 9 / Number of copies: 1 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.330 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 57.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 0.0 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 64000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)