[English] 日本語
 Yorodumi
Yorodumi- EMDB-40918: Human glutaminase C (Y466W) with L-Gln and Pi, filamentous form -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human glutaminase C (Y466W) with L-Gln and Pi, filamentous form | ||||||||||||||||||||||||
 Map data Map data | |||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||
 Keywords Keywords | Cancer / Filament / Metabolism / HYDROLASE | ||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationL-glutamine catabolic process / regulation of respiratory gaseous exchange by nervous system process / glutaminase / intracellular glutamate homeostasis / Glutamate and glutamine metabolism / Glutamate Neurotransmitter Release Cycle / : / glutaminase activity / suckling behavior / TP53 Regulates Metabolic Genes ...L-glutamine catabolic process / regulation of respiratory gaseous exchange by nervous system process / glutaminase / intracellular glutamate homeostasis / Glutamate and glutamine metabolism / Glutamate Neurotransmitter Release Cycle / : / glutaminase activity / suckling behavior / TP53 Regulates Metabolic Genes / protein homotetramerization / chemical synaptic transmission / mitochondrial matrix / synapse / mitochondrion / cytosol Similarity search - Function | ||||||||||||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||||||||||||||
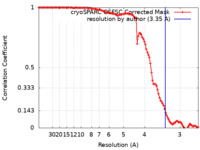
| Method | helical reconstruction / cryo EM / Resolution: 3.35 Å | ||||||||||||||||||||||||
 Authors Authors | Feng S / Aplin C / Nguyen T-TT / Milano SK / Cerione RA | ||||||||||||||||||||||||
| Funding support |  United States, 7 items United States, 7 items
| ||||||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Filament formation drives catalysis by glutaminase enzymes important in cancer progression. Authors: Shi Feng / Cody Aplin / Thuy-Tien T Nguyen / Shawn K Milano / Richard A Cerione /  Abstract: The glutaminase enzymes GAC and GLS2 catalyze the hydrolysis of glutamine to glutamate, satisfying the 'glutamine addiction' of cancer cells. They are the targets of anti-cancer drugs; however, their ...The glutaminase enzymes GAC and GLS2 catalyze the hydrolysis of glutamine to glutamate, satisfying the 'glutamine addiction' of cancer cells. They are the targets of anti-cancer drugs; however, their mechanisms of activation and catalytic activity have been unclear. Here we demonstrate that the ability of GAC and GLS2 to form filaments is directly coupled to their catalytic activity and present their cryo-EM structures which provide a view of the conformational states essential for catalysis. Filament formation guides an 'activation loop' to assume a specific conformation that works together with a 'lid' to close over the active site and position glutamine for nucleophilic attack by an essential serine. Our findings highlight how ankyrin repeats on GLS2 regulate enzymatic activity, while allosteric activators stabilize, and clinically relevant inhibitors block, filament formation that enables glutaminases to catalyze glutaminolysis and support cancer progression. | ||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40918.map.gz emd_40918.map.gz | 117.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40918-v30.xml emd-40918-v30.xml emd-40918.xml emd-40918.xml | 15.8 KB 15.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40918_fsc.xml emd_40918_fsc.xml | 10.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_40918.png emd_40918.png | 76.9 KB | ||
| Filedesc metadata |  emd-40918.cif.gz emd-40918.cif.gz | 5.9 KB | ||
| Others |  emd_40918_half_map_1.map.gz emd_40918_half_map_1.map.gz emd_40918_half_map_2.map.gz emd_40918_half_map_2.map.gz | 115.8 MB 115.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40918 http://ftp.pdbj.org/pub/emdb/structures/EMD-40918 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40918 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40918 | HTTPS FTP |
-Related structure data
| Related structure data |  8szjMC  8szlC  8t0zC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40918.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40918.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.31 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_40918_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
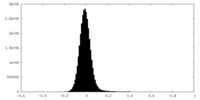
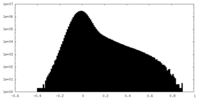
| Density Histograms |
-Half map: #1
| File | emd_40918_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
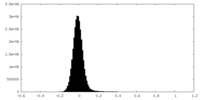
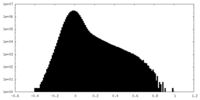
| Density Histograms |
- Sample components
Sample components
-Entire : Human glutaminase C (Y466W) with L-Gln and Pi
| Entire | Name: Human glutaminase C (Y466W) with L-Gln and Pi |
|---|---|
| Components |
|
-Supramolecule #1: Human glutaminase C (Y466W) with L-Gln and Pi
| Supramolecule | Name: Human glutaminase C (Y466W) with L-Gln and Pi / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Glutaminase kidney isoform, mitochondrial
| Macromolecule | Name: Glutaminase kidney isoform, mitochondrial / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO / EC number: glutaminase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 65.563953 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MMRLRGSGML RDLLLRSPAG VSATLRRAQP LVTLCRRPRG GGRPAAGPAA AARLHPWWGG GGWPAEPLAR GLSSSPSEIL QELGKGSTH PQPGVSPPAA PAAPGPKDGP GETDAFGNSE GKELVASGEN KIKQGLLPSL EDLLFYTIAE GQEKIPVHKF I TALKSTGL ...String: MMRLRGSGML RDLLLRSPAG VSATLRRAQP LVTLCRRPRG GGRPAAGPAA AARLHPWWGG GGWPAEPLAR GLSSSPSEIL QELGKGSTH PQPGVSPPAA PAAPGPKDGP GETDAFGNSE GKELVASGEN KIKQGLLPSL EDLLFYTIAE GQEKIPVHKF I TALKSTGL RTSDPRLKEC MDMLRLTLQT TSDGVMLDKD LFKKCVQSNI VLLTQAFRRK FVIPDFMSFT SHIDELYESA KK QSGGKVA DYIPQLAKFS PDLWGVSVCT VDGQRHSTGD TKVPFCLQSC VKPLKYAIAV NDLGTEYVHR YVGKEPSGLR FNK LFLNED DKPHNPMVNA GAIVVTSLIK QGVNNAEKFD YVMQFLNKMA GNEYVGFSNA TFQSERESGD RNFAIGYYLK EKKC FPEGT DMVGILDFYF QLCSIEVTCE SASVMAATLA NGGFCPITGE RVLSPEAVRN TLSLMHSCGM WDFSGQFAFH VGLPA KSGV AGGILLVVPN VMGMMCWSPP LDKMGNSVKG IHFCHDLVSL CNFHNYDNLR HFAKKLDPRR EGGDQRHSFG PLDYES LQQ ELALKETVWK KVSPESNEDI STTVVYRMES LGEKS UniProtKB: Glutaminase kidney isoform, mitochondrial |
-Macromolecule #2: PHOSPHATE ION
| Macromolecule | Name: PHOSPHATE ION / type: ligand / ID: 2 / Number of copies: 12 / Formula: PO4 |
|---|---|
| Molecular weight | Theoretical: 94.971 Da |
| Chemical component information |  ChemComp-PO4: |
-Macromolecule #3: GLUTAMINE
| Macromolecule | Name: GLUTAMINE / type: ligand / ID: 3 / Number of copies: 12 / Formula: GLN |
|---|---|
| Molecular weight | Theoretical: 146.144 Da |
| Chemical component information | 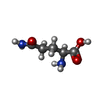 ChemComp-GLN: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)