+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | TMEM16F bound with Niclosamide | |||||||||
 Map data Map data | TMEM16F Niclosamide | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ca2+-activated ion channels and lipid scramblases / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcalcium activated galactosylceramide scrambling / phosphatidylserine exposure on blood platelet / calcium activated phosphatidylserine scrambling / calcium activated phosphatidylcholine scrambling / calcium activated phospholipid scrambling / positive regulation of potassium ion export across plasma membrane / positive regulation of monoatomic ion transmembrane transport / activation of blood coagulation via clotting cascade / purinergic nucleotide receptor signaling pathway / phospholipid scramblase activity ...calcium activated galactosylceramide scrambling / phosphatidylserine exposure on blood platelet / calcium activated phosphatidylserine scrambling / calcium activated phosphatidylcholine scrambling / calcium activated phospholipid scrambling / positive regulation of potassium ion export across plasma membrane / positive regulation of monoatomic ion transmembrane transport / activation of blood coagulation via clotting cascade / purinergic nucleotide receptor signaling pathway / phospholipid scramblase activity / bone mineralization involved in bone maturation / cholinergic synapse / intracellularly calcium-gated chloride channel activity / pore complex assembly / negative regulation of cell volume / plasma membrane phospholipid scrambling / voltage-gated monoatomic ion channel activity / bleb assembly / Stimuli-sensing channels / positive regulation of phagocytosis, engulfment / voltage-gated chloride channel activity / calcium-activated cation channel activity / positive regulation of monocyte chemotaxis / dendritic cell chemotaxis / chloride transport / phospholipid translocation / chloride channel activity / regulation of postsynaptic membrane potential / positive regulation of endothelial cell apoptotic process / positive regulation of bone mineralization / chloride channel complex / Neutrophil degranulation / chloride transmembrane transport / sodium ion transmembrane transport / synaptic membrane / calcium ion transmembrane transport / blood coagulation / positive regulation of apoptotic process / protein homodimerization activity / metal ion binding / identical protein binding / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / Resolution: 3.1 Å | |||||||||
 Authors Authors | Feng S / Cheng Y | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Identification of a drug binding pocket in TMEM16F calcium-activated ion channel and lipid scramblase. Authors: Shengjie Feng / Cristina Puchades / Juyeon Ko / Hao Wu / Yifei Chen / Eric E Figueroa / Shuo Gu / Tina W Han / Brandon Ho / Tong Cheng / Junrui Li / Brian Shoichet / Yuh Nung Jan / Yifan Cheng / Lily Yeh Jan /  Abstract: The dual functions of TMEM16F as Ca-activated ion channel and lipid scramblase raise intriguing questions regarding their molecular basis. Intrigued by the ability of the FDA-approved drug ...The dual functions of TMEM16F as Ca-activated ion channel and lipid scramblase raise intriguing questions regarding their molecular basis. Intrigued by the ability of the FDA-approved drug niclosamide to inhibit TMEM16F-dependent syncytia formation induced by SARS-CoV-2, we examined cryo-EM structures of TMEM16F with or without bound niclosamide or 1PBC, a known blocker of TMEM16A Ca-activated Cl channel. Here, we report evidence for a lipid scrambling pathway along a groove harboring a lipid trail outside the ion permeation pore. This groove contains the binding pocket for niclosamide and 1PBC. Mutations of two residues in this groove specifically affect lipid scrambling. Whereas mutations of some residues in the binding pocket of niclosamide and 1PBC reduce their inhibition of TMEM16F-mediated Ca influx and PS exposure, other mutations preferentially affect the ability of niclosamide and/or 1PBC to inhibit TMEM16F-mediated PS exposure, providing further support for separate pathways for ion permeation and lipid scrambling. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40776.map.gz emd_40776.map.gz | 59.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40776-v30.xml emd-40776-v30.xml emd-40776.xml emd-40776.xml | 15.2 KB 15.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_40776.png emd_40776.png | 66.3 KB | ||
| Filedesc metadata |  emd-40776.cif.gz emd-40776.cif.gz | 6 KB | ||
| Others |  emd_40776_half_map_1.map.gz emd_40776_half_map_1.map.gz emd_40776_half_map_2.map.gz emd_40776_half_map_2.map.gz | 11.7 MB 11.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40776 http://ftp.pdbj.org/pub/emdb/structures/EMD-40776 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40776 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40776 | HTTPS FTP |
-Related structure data
| Related structure data |  8surMC  8sunC  8tagC  8taiC  8talC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40776.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40776.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | TMEM16F Niclosamide | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.839 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: TMEM16F Niclosamide halfmap1
| File | emd_40776_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | TMEM16F Niclosamide halfmap1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: TMEM16F Niclosamide halfmap2
| File | emd_40776_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | TMEM16F Niclosamide halfmap2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : 16F
| Entire | Name: 16F |
|---|---|
| Components |
|
-Supramolecule #1: 16F
| Supramolecule | Name: 16F / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Anoctamin-6
| Macromolecule | Name: Anoctamin-6 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 106.367727 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MQMMTRKVLL NMELEEDDDE DGDIVLENFD QTIVCPTFGS LENQQDFRTP EFEEFNGKPD SLFFTDGQRR IDFILVYEDE SKKENNKKG TNEKQKRKRQ AYESNLICHG LQLEATRSVS DDKLVFVKVH APWEVLCTYA EIMHIKLPLK PNDLKTRSPF G NLNWFTKV ...String: MQMMTRKVLL NMELEEDDDE DGDIVLENFD QTIVCPTFGS LENQQDFRTP EFEEFNGKPD SLFFTDGQRR IDFILVYEDE SKKENNKKG TNEKQKRKRQ AYESNLICHG LQLEATRSVS DDKLVFVKVH APWEVLCTYA EIMHIKLPLK PNDLKTRSPF G NLNWFTKV LRVNESVIKP EQEFFTAPFE KSRMNDFYIL DRDSFFNPAT RSRIVYFILS RVKYQVMNNV NKFGINRLVS SG IYKAAFP LHDCRFNYES EDISCPSERY LLYREWAHPR SIYKKQPLDL IRKYYGEKIG IYFAWLGYYT QMLLLAAVVG VAC FLYGYL DQDNCTWSKE VCDPDIGGQI LMCPQCDRLC PFWRLNITCE SSKKLCIFDS FGTLIFAVFM GVWVTLFLEF WKRR QAELE YEWDTVELQQ EEQARPEYEA QCNHVVINEI TQEEERIPFT TCGKCIRVTL CASAVFFWIL LIIASVIGII VYRLS VFIV FSTTLPKNPN GTDPIQKYLT PQMATSITAS IISFIIIMIL NTIYEKVAIM ITNFELPRTQ TDYENSLTMK MFLFQF VNY YSSCFYIAFF KGKFVGYPGD PVYLLGKYRS EECDPGGCLL ELTTQLTIIM GGKAIWNNIQ EVLLPWVMNL IGRYKRV SG SEKITPRWEQ DYHLQPMGKL GLFYEYLEMI IQFGFVTLFV ASFPLAPLLA LVNNILEIRV DAWKLTTQFR RMVPEKAQ D IGAWQPIMQG IAILAVVTNA MIIAFTSDMI PRLVYYWSFS IPPYGDHTYY TMDGYINNTL SVFNITDFKN TDKENPYIG LGNYTLCRYR DFRNPPGHPQ EYKHNIYYWH VIAAKLAFII VMEHIIYSVK FFISYAIPDV SKITKSKIKR EKYLTQKLLH ESHLKDLTK NMGIIAERIG GTVDNSVRPK LE UniProtKB: Anoctamin-6 |
-Macromolecule #3: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 3 / Number of copies: 5 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 3 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #5: 5-chloro-N-(2-chloro-4-nitrophenyl)-2-hydroxybenzamide
| Macromolecule | Name: 5-chloro-N-(2-chloro-4-nitrophenyl)-2-hydroxybenzamide type: ligand / ID: 5 / Number of copies: 1 / Formula: VUT |
|---|---|
| Molecular weight | Theoretical: 327.12 Da |
| Chemical component information | 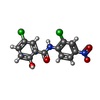 ChemComp-VUT: |
-Experimental details
-Structure determination
 Processing Processing | single particle reconstruction |
|---|---|
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 66.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 1047816 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)