+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4050 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | 70S ribosome from Staphylococcus aureus | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | pathogenic ribosome / bL31 B-type / ribosome | |||||||||
| Function / homology |  Function and homology information Function and homology informationlarge ribosomal subunit / transferase activity / ribosome biogenesis / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding ...large ribosomal subunit / transferase activity / ribosome biogenesis / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / ribonucleoprotein complex / mRNA binding / RNA binding / zinc ion binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Staphylococcus aureus (strain NCTC 8325) (bacteria) Staphylococcus aureus (strain NCTC 8325) (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Khusainov I / Vicens Q | |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2016 Journal: Nucleic Acids Res / Year: 2016Title: Structure of the 70S ribosome from human pathogen Staphylococcus aureus. Authors: Iskander Khusainov / Quentin Vicens / Anthony Bochler / François Grosse / Alexander Myasnikov / Jean-François Ménétret / Johana Chicher / Stefano Marzi / Pascale Romby / Gulnara Yusupova ...Authors: Iskander Khusainov / Quentin Vicens / Anthony Bochler / François Grosse / Alexander Myasnikov / Jean-François Ménétret / Johana Chicher / Stefano Marzi / Pascale Romby / Gulnara Yusupova / Marat Yusupov / Yaser Hashem /   Abstract: Comparative structural studies of ribosomes from various organisms keep offering exciting insights on how species-specific or environment-related structural features of ribosomes may impact ...Comparative structural studies of ribosomes from various organisms keep offering exciting insights on how species-specific or environment-related structural features of ribosomes may impact translation specificity and its regulation. Although the importance of such features may be less obvious within more closely related organisms, their existence could account for vital yet species-specific mechanisms of translation regulation that would involve stalling, cell survival and antibiotic resistance. Here, we present the first full 70S ribosome structure from Staphylococcus aureus, a Gram-positive pathogenic bacterium, solved by cryo-electron microscopy. Comparative analysis with other known bacterial ribosomes pinpoints several unique features specific to S. aureus around a conserved core, at both the protein and the RNA levels. Our work provides the structural basis for the many studies aiming at understanding translation regulation in S. aureus and for designing drugs against this often multi-resistant pathogen. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4050.map.gz emd_4050.map.gz | 130.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4050-v30.xml emd-4050-v30.xml emd-4050.xml emd-4050.xml | 64.4 KB 64.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_4050.jpg emd_4050.jpg emd_4050.png emd_4050.png | 589.3 KB 332.7 KB | ||
| Filedesc metadata |  emd-4050.cif.gz emd-4050.cif.gz | 13 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4050 http://ftp.pdbj.org/pub/emdb/structures/EMD-4050 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4050 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4050 | HTTPS FTP |
-Related structure data
| Related structure data |  5li0MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4050.map.gz / Format: CCP4 / Size: 139.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4050.map.gz / Format: CCP4 / Size: 139.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
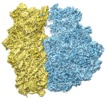
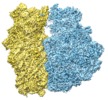
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : 70S ribosome from Staphylococcus aureus
+Supramolecule #1: 70S ribosome from Staphylococcus aureus
+Macromolecule #1: 16S ribosomal RNA
+Macromolecule #21: 23S ribosomal RNA
+Macromolecule #22: 5S ribosomal RNA
+Macromolecule #2: 30S ribosomal protein S2
+Macromolecule #3: 30S ribosomal protein S3
+Macromolecule #4: 30S ribosomal protein S4
+Macromolecule #5: 30S ribosomal protein S5
+Macromolecule #6: 30S ribosomal protein S6
+Macromolecule #7: 30S ribosomal protein S7
+Macromolecule #8: 30S ribosomal protein S8
+Macromolecule #9: 30S ribosomal protein S9
+Macromolecule #10: 30S ribosomal protein S10
+Macromolecule #11: 30S ribosomal protein S11
+Macromolecule #12: 30S ribosomal protein S12
+Macromolecule #13: 30S ribosomal protein S13
+Macromolecule #14: 30S ribosomal protein S14 type Z
+Macromolecule #15: 30S ribosomal protein S15
+Macromolecule #16: 30S ribosomal protein S16
+Macromolecule #17: 30S ribosomal protein S17
+Macromolecule #18: 30S ribosomal protein S18
+Macromolecule #19: 30S ribosomal protein S19
+Macromolecule #20: 30S ribosomal protein S20
+Macromolecule #23: 50S ribosomal protein L2
+Macromolecule #24: 50S ribosomal protein L3
+Macromolecule #25: 50S ribosomal protein L4
+Macromolecule #26: 50S ribosomal protein L5
+Macromolecule #27: 50S ribosomal protein L6
+Macromolecule #28: 50S ribosomal protein L13
+Macromolecule #29: 50S ribosomal protein L14
+Macromolecule #30: 50S ribosomal protein L15
+Macromolecule #31: 50S ribosomal protein L16
+Macromolecule #32: 50S ribosomal protein L17
+Macromolecule #33: 50S ribosomal protein L18
+Macromolecule #34: 50S ribosomal protein L19
+Macromolecule #35: 50S ribosomal protein L20
+Macromolecule #36: 50S ribosomal protein L21
+Macromolecule #37: 50S ribosomal protein L22
+Macromolecule #38: 50S ribosomal protein L23
+Macromolecule #39: 50S ribosomal protein L24
+Macromolecule #40: 50S ribosomal protein L25
+Macromolecule #41: 50S ribosomal protein L27
+Macromolecule #42: 50S ribosomal protein L28
+Macromolecule #43: 50S ribosomal protein L29
+Macromolecule #44: 50S ribosomal protein L30
+Macromolecule #45: 50S ribosomal protein L31 type B
+Macromolecule #46: 50S ribosomal protein L32
+Macromolecule #47: 50S ribosomal protein L33 2
+Macromolecule #48: 50S ribosomal protein L34
+Macromolecule #49: 50S ribosomal protein L35
+Macromolecule #50: 50S ribosomal protein L36
+Macromolecule #51: MAGNESIUM ION
+Macromolecule #52: OXYGEN ATOM
+Macromolecule #53: ZINC ION
+Macromolecule #54: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Details: 5 mM Hepes-KOH pH 7.5 50 mM KCl 10 mM NH4Cl 10 mM Mg(OAc)2 1 mM DTT |
|---|---|
| Grid | Material: COPPER / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: blot force 5, blot waiting time 30 s. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 2-8 / Number grids imaged: 1 / Number real images: 3850 / Average exposure time: 1.0 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: DARK FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER |
|---|---|
| Output model |  PDB-5li0: |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)
























