+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of P-Glycoprotein in Apo state | |||||||||
 Map data Map data | Pgp in lipid nanodiscs in Apo form | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | multidrug resistance / ABC transporter / membrane protein / transporter / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationhormone transport / phosphatidylethanolamine floppase activity / cellular response to nonylphenol / cellular response to borneol / response to codeine / cellular response to mycotoxin / daunorubicin transport / positive regulation of response to drug / terpenoid transport / ceramide floppase activity ...hormone transport / phosphatidylethanolamine floppase activity / cellular response to nonylphenol / cellular response to borneol / response to codeine / cellular response to mycotoxin / daunorubicin transport / positive regulation of response to drug / terpenoid transport / ceramide floppase activity / regulation of intestinal absorption / cellular response to external biotic stimulus / response to cyclosporin A / response to antineoplastic agent / positive regulation of establishment of Sertoli cell barrier / negative regulation of sensory perception of pain / carboxylic acid transmembrane transport / floppase activity / ceramide translocation / Abacavir transmembrane transport / response to quercetin / carboxylic acid transmembrane transporter activity / establishment of blood-retinal barrier / phosphatidylethanolamine flippase activity / protein localization to bicellular tight junction / phosphatidylcholine floppase activity / external side of apical plasma membrane / Atorvastatin ADME / response to thyroxine / xenobiotic transport across blood-brain barrier / establishment of blood-brain barrier / export across plasma membrane / P-type phospholipid transporter / xenobiotic detoxification by transmembrane export across the plasma membrane / transepithelial transport / ABC-type xenobiotic transporter / cellular response to L-glutamate / response to vitamin A / response to glucagon / response to vitamin D / intestinal absorption / response to alcohol / response to glycoside / Prednisone ADME / ABC-type xenobiotic transporter activity / phospholipid translocation / cellular hyperosmotic salinity response / cellular response to alkaloid / cellular response to antibiotic / maintenance of blood-brain barrier / ATPase-coupled transmembrane transporter activity / efflux transmembrane transporter activity / xenobiotic transmembrane transporter activity / cellular response to dexamethasone stimulus / response to cadmium ion / transmembrane transporter activity / transport across blood-brain barrier / lactation / response to progesterone / xenobiotic metabolic process / regulation of chloride transport / placenta development / stem cell proliferation / cellular response to estradiol stimulus / brush border membrane / female pregnancy / circadian rhythm / transmembrane transport / ABC-family protein mediated transport / G2/M transition of mitotic cell cycle / cellular response to tumor necrosis factor / cellular response to lipopolysaccharide / response to hypoxia / apical plasma membrane / response to xenobiotic stimulus / ubiquitin protein ligase binding / cell surface / ATP hydrolysis activity / extracellular exosome / ATP binding / membrane / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.3 Å | |||||||||
 Authors Authors | Culbertson A / Liao M | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2025 Journal: Nat Commun / Year: 2025Title: Cryo-EM of human P-glycoprotein reveals an intermediate occluded conformation during active drug transport. Authors: Alan T Culbertson / Maofu Liao /   Abstract: P-glycoprotein (Pgp) is an important human multidrug transporter that contributes to pharmacokinetics and multidrug resistance. Despite decades of study, the conformation transition cycle of Pgp ...P-glycoprotein (Pgp) is an important human multidrug transporter that contributes to pharmacokinetics and multidrug resistance. Despite decades of study, the conformation transition cycle of Pgp undergoing active drug transport is not defined, thus the precise relevance of all available Pgp structures to uninterrupted multidrug transport remains unclear. Here, we use cryo-EM of membrane-embedded human Pgp under continuous turnover conditions to analyze the conformational ensembles of Pgp transporting distinct substrates. These results delineate multiple conformations including inward-facing and closed conformations, highlighting the occluded conformation as a critical intermediate state between transporter closure and substrate release. A combination of structural, functional, and computational studies reveals the transmembrane helices 4 and 10 undergoing drastic rearrangement to coordinate substrate binding, occlusion, and release, and identifies a peripheral site involved in substrate capture and Pgp inhibition. Together, our results provide a set of snapshots of Pgp undergoing continuous drug transport, unveiling the intricate interplay between transporter dynamics and drug movement, and shed light on the mechanism of polyspecificity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40226.map.gz emd_40226.map.gz | 24.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40226-v30.xml emd-40226-v30.xml emd-40226.xml emd-40226.xml | 18.1 KB 18.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_40226.png emd_40226.png | 111.6 KB | ||
| Filedesc metadata |  emd-40226.cif.gz emd-40226.cif.gz | 6.5 KB | ||
| Others |  emd_40226_half_map_1.map.gz emd_40226_half_map_1.map.gz emd_40226_half_map_2.map.gz emd_40226_half_map_2.map.gz | 20.6 MB 20.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40226 http://ftp.pdbj.org/pub/emdb/structures/EMD-40226 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40226 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40226 | HTTPS FTP |
-Related structure data
| Related structure data |  8gmgMC  8gmjC  8sa0C  8sa1C  8sb7C  8sb8C  8sb9C  8sbaC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40226.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40226.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Pgp in lipid nanodiscs in Apo form | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
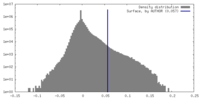
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Pgp in lipid nanodiscs in Apo form
| File | emd_40226_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Pgp in lipid nanodiscs in Apo form | ||||||||||||
| Projections & Slices |
| ||||||||||||
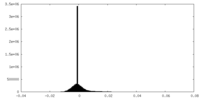
| Density Histograms |
-Half map: Pgp in lipid nanodiscs in Apo form
| File | emd_40226_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Pgp in lipid nanodiscs in Apo form | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CryoEM structure of P-Glycoprotein in Apo state
| Entire | Name: CryoEM structure of P-Glycoprotein in Apo state |
|---|---|
| Components |
|
-Supramolecule #1: CryoEM structure of P-Glycoprotein in Apo state
| Supramolecule | Name: CryoEM structure of P-Glycoprotein in Apo state / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: ATP-dependent translocase ABCB1
| Macromolecule | Name: ATP-dependent translocase ABCB1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: ABC-type xenobiotic transporter |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 141.644781 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDLEGDRNGG AKKKNFFKLN NKSEKDKKEK KPTVSVFSMF RYSNWLDKLY MVVGTLAAII HGAGLPLMML VFGEMTDIFA NAGNLEDLM SNITNRSDIN DTGFFMNLEE DMTRYAYYYS GIGAGVLVAA YIQVSFWCLA AGRQIHKIRK QFFHAIMRQE I GWFDVHDV ...String: MDLEGDRNGG AKKKNFFKLN NKSEKDKKEK KPTVSVFSMF RYSNWLDKLY MVVGTLAAII HGAGLPLMML VFGEMTDIFA NAGNLEDLM SNITNRSDIN DTGFFMNLEE DMTRYAYYYS GIGAGVLVAA YIQVSFWCLA AGRQIHKIRK QFFHAIMRQE I GWFDVHDV GELNTRLTDD VSKINEGIGD KIGMFFQSMA TFFTGFIVGF TRGWKLTLVI LAISPVLGLS AAVWAKILSS FT DKELLAY AKAGAVAEEV LAAIRTVIAF GGQKKELERY NKNLEEAKRI GIKKAITANI SIGAAFLLIY ASYALAFWYG TTL VLSGEY SIGQVLTVFF SVLIGAFSVG QASPSIEAFA NARGAAYEIF KIIDNKPSID SYSKSGHKPD NIKGNLEFRN VHFS YPSRK EVKILKGLNL KVQSGQTVAL VGNSGCGKST TVQLMQRLYD PTEGMVSVDG QDIRTINVRF LREIIGVVSQ EPVLF ATTI AENIRYGREN VTMDEIEKAV KEANAYDFIM KLPHKFDTLV GERGAQLSGG QKQRIAIARA LVRNPKILLL DEATSA LDT ESEAVVQVAL DKARKGRTTI VIAHRLSTVR NADVIAGFDD GVIVEKGNHD ELMKEKGIYF KLVTMQTAGN EVELENA AD ESKSEIDALE MSSNDSRSSL IRKRSTRRSV RGSQAQDRKL STKEALDESI PPVSFWRIMK LNLTEWPYFV VGVFCAII N GGLQPAFAII FSKIIGVFTR IDDPETKRQN SNLFSLLFLA LGIISFITFF LQGFTFGKAG EILTKRLRYM VFRSMLRQD VSWFDDPKNT TGALTTRLAN DAAQVKGAIG SRLAVITQNI ANLGTGIIIS FIYGWQLTLL LLAIVPIIAI AGVVEMKMLS GQALKDKKE LEGSGKIATE AIENFRTVVS LTQEQKFEHM YAQSLQVPYR NSLRKAHIFG ITFSFTQAMM YFSYAGCFRF G AYLVAHKL MSFEDVLLVF SAVVFGAMAV GQVSSFAPDY AKAKISAAHI IMIIEKTPLI DSYSTEGLMP NTLEGNVTFG EV VFNYPTR PDIPVLQGLS LEVKKGQTLA LVGSSGCGKS TVVQLLERFY DPLAGKVLLD GKEIKRLNVQ WLRAHLGIVS QEP ILFDCS IAENIAYGDN SRVVSQEEIV RAAKEANIHA FIESLPNKYS TKVGDKGTQL SGGQKQRIAI ARALVRQPHI LLLD EATSA LDTESEKVVQ EALDKAREGR TCIVIAHRLS TIQNADLIVV FQNGRVKEHG THQQLLAQKG IYFSMVSVQA GTKRQ UniProtKB: ATP-dependent translocase ABCB1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 2 / Number real images: 8983 / Average exposure time: 3.5 sec. / Average electron dose: 51.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 81000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)