+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3925 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
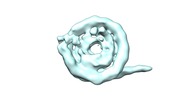
| Title | Nucleosome breathing : Class 9 | |||||||||
 Map data Map data | Nucleosome breathing Class 9 | |||||||||
 Sample Sample |
| |||||||||
| Biological species | ||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 8.3 Å | |||||||||
 Authors Authors | Halic M / Bilokapic S | |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2018 Journal: Nat Struct Mol Biol / Year: 2018Title: Histone octamer rearranges to adapt to DNA unwrapping. Authors: Silvija Bilokapic / Mike Strauss / Mario Halic /  Abstract: Nucleosomes, the basic units of chromatin, package and regulate expression of eukaryotic genomes. Although the structure of the intact nucleosome is well characterized, little is known about ...Nucleosomes, the basic units of chromatin, package and regulate expression of eukaryotic genomes. Although the structure of the intact nucleosome is well characterized, little is known about structures of partially unwrapped, transient intermediates. In this study, we present nine cryo-EM structures of distinct conformations of nucleosome and subnucleosome particles. These structures show that initial DNA breathing induces conformational changes in the histone octamer, particularly in histone H3, that propagate through the nucleosome and prevent symmetrical DNA opening. Rearrangements in the H2A-H2B dimer strengthen interaction with the unwrapping DNA and promote nucleosome stability. In agreement with this, cross-linked H2A-H2B that cannot accommodate unwrapping of the DNA is not stably maintained in the nucleosome. H2A-H2B release and DNA unwrapping occur simultaneously, indicating that DNA is essential in stabilizing the dimer in the nucleosome. Our structures reveal intrinsic nucleosomal plasticity that is required for nucleosome stability and might be exploited by extrinsic protein factors. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3925.map.gz emd_3925.map.gz | 11.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3925-v30.xml emd-3925-v30.xml emd-3925.xml emd-3925.xml | 7.7 KB 7.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_3925.png emd_3925.png | 137.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3925 http://ftp.pdbj.org/pub/emdb/structures/EMD-3925 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3925 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3925 | HTTPS FTP |
-Related structure data
| Related structure data |  3926C  3929C  3930C  3931C  3947C  3948C  3949C  3950C  6esfC  6esgC  6eshC  6esiC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3925.map.gz / Format: CCP4 / Size: 18.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3925.map.gz / Format: CCP4 / Size: 18.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Nucleosome breathing Class 9 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.4 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Nucleosome
| Entire | Name: Nucleosome |
|---|---|
| Components |
|
-Supramolecule #1: Nucleosome
| Supramolecule | Name: Nucleosome / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism: |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 200 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Average electron dose: 100.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 8.3 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 1.4) / Number images used: 9000 |
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






















