+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of ACPII-CCPII from cryptophyte algae | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Photosystem II / PHOTOSYNTHESIS / STRUCTURAL PROTEIN | |||||||||
| Biological species |  Chroomonas placoidea (eukaryote) Chroomonas placoidea (eukaryote) | |||||||||
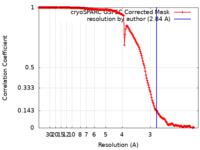
| Method | single particle reconstruction / cryo EM / Resolution: 2.84 Å | |||||||||
 Authors Authors | Li XY / Mao ZY / Shen JR / Han GY | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structure and distinct supramolecular organization of a PSII-ACPII dimer from a cryptophyte alga Chroomonas placoidea. Authors: Zhiyuan Mao / Xingyue Li / Zhenhua Li / Liangliang Shen / Xiaoyi Li / Yanyan Yang / Wenda Wang / Tingyun Kuang / Jian-Ren Shen / Guangye Han /   Abstract: Cryptophyte algae are an evolutionarily distinct and ecologically important group of photosynthetic unicellular eukaryotes. Photosystem II (PSII) of cryptophyte algae associates with alloxanthin ...Cryptophyte algae are an evolutionarily distinct and ecologically important group of photosynthetic unicellular eukaryotes. Photosystem II (PSII) of cryptophyte algae associates with alloxanthin chlorophyll a/c-binding proteins (ACPs) to act as the peripheral light-harvesting system, whose supramolecular organization is unknown. Here, we purify the PSII-ACPII supercomplex from a cryptophyte alga Chroomonas placoidea (C. placoidea), and analyze its structure at a resolution of 2.47 Å using cryo-electron microscopy. This structure reveals a dimeric organization of PSII-ACPII containing two PSII core monomers flanked by six symmetrically arranged ACPII subunits. The PSII core is conserved whereas the organization of ACPII subunits exhibits a distinct pattern, different from those observed so far in PSII of other algae and higher plants. Furthermore, we find a Chl a-binding antenna subunit, CCPII-S, which mediates interaction of ACPII with the PSII core. These results provide a structural basis for the assembly of antennas within the supercomplex and possible excitation energy transfer pathways in cryptophyte algal PSII, shedding light on the diversity of supramolecular organization of photosynthetic machinery. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38419.map.gz emd_38419.map.gz | 256.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38419-v30.xml emd-38419-v30.xml emd-38419.xml emd-38419.xml | 23.7 KB 23.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_38419_fsc.xml emd_38419_fsc.xml | 16.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_38419.png emd_38419.png | 61.1 KB | ||
| Filedesc metadata |  emd-38419.cif.gz emd-38419.cif.gz | 6.9 KB | ||
| Others |  emd_38419_half_map_1.map.gz emd_38419_half_map_1.map.gz emd_38419_half_map_2.map.gz emd_38419_half_map_2.map.gz | 474.4 MB 474.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38419 http://ftp.pdbj.org/pub/emdb/structures/EMD-38419 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38419 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38419 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_38419.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38419.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
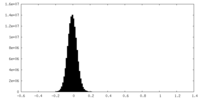
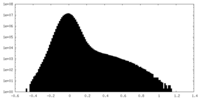
-Half map: #2
| File | emd_38419_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
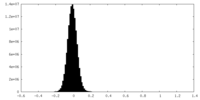
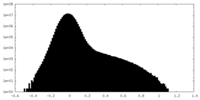
| Density Histograms |
-Half map: #1
| File | emd_38419_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Structure of ACPII-CCPII from cryptophyte algae
+Supramolecule #1: Structure of ACPII-CCPII from cryptophyte algae
+Macromolecule #1: ACPII-4
+Macromolecule #2: ACPII-1
+Macromolecule #3: ACPII-2
+Macromolecule #4: ACPII-3
+Macromolecule #5: Photosystem II reaction center protein G
+Macromolecule #6: ACPII-6
+Macromolecule #7: ACPII-5
+Macromolecule #8: CCPII-S
+Macromolecule #9: CHLOROPHYLL A
+Macromolecule #10: Chlorophyll c2
+Macromolecule #11: (1~{R})-3,5,5-trimethyl-4-[(3~{E},5~{E},7~{E},9~{E},11~{E},13~{E}...
+Macromolecule #12: (1~{R})-3,5,5-trimethyl-4-[(3~{E},5~{E},7~{E},9~{E},11~{E},13~{E}...
+Macromolecule #13: 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE
+Macromolecule #14: (1~{R})-3,5,5-trimethyl-4-[(3~{E},5~{E},7~{E},9~{E},11~{E},13~{E}...
+Macromolecule #15: 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE
+Macromolecule #16: (6'R,11cis,11'cis,13cis,15cis)-4',5'-didehydro-5',6'-dihydro-beta...
+Macromolecule #17: 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)