[English] 日本語
 Yorodumi
Yorodumi- EMDB-38232: The cryo-EM structure of the RAD51 L2 loop bound to the linker DN... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | The cryo-EM structure of the RAD51 L2 loop bound to the linker DNA with the blunt end of the nucleosome | ||||||||||||||||||
 Map data Map data | |||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Nucleosome / Recombinase / DNA BINDING PROTEIN-DNA Complex | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationpresynaptic intermediate filament cytoskeleton / response to glucoside / mitotic recombination-dependent replication fork processing / DNA recombinase assembly / chromosome organization involved in meiotic cell cycle / telomere maintenance via telomere lengthening / double-strand break repair involved in meiotic recombination / nuclear ubiquitin ligase complex / cellular response to cisplatin / DNA strand invasion ...presynaptic intermediate filament cytoskeleton / response to glucoside / mitotic recombination-dependent replication fork processing / DNA recombinase assembly / chromosome organization involved in meiotic cell cycle / telomere maintenance via telomere lengthening / double-strand break repair involved in meiotic recombination / nuclear ubiquitin ligase complex / cellular response to cisplatin / DNA strand invasion / cellular response to hydroxyurea / mitotic recombination / cellular response to camptothecin / replication-born double-strand break repair via sister chromatid exchange / lateral element / regulation of DNA damage checkpoint / DNA strand exchange activity / Impaired BRCA2 binding to PALB2 / telomere maintenance via recombination / single-stranded DNA helicase activity / reciprocal meiotic recombination / Homologous DNA Pairing and Strand Exchange / Defective homologous recombination repair (HRR) due to BRCA1 loss of function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA1 binding function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA2/RAD51/RAD51C binding function / Resolution of D-loop Structures through Synthesis-Dependent Strand Annealing (SDSA) / Resolution of D-loop Structures through Holliday Junction Intermediates / HDR through Single Strand Annealing (SSA) / ATP-dependent DNA damage sensor activity / regulation of double-strand break repair via homologous recombination / nuclear chromosome / Impaired BRCA2 binding to RAD51 / Transcriptional Regulation by E2F6 / replication fork processing / Presynaptic phase of homologous DNA pairing and strand exchange / response to X-ray / ATP-dependent activity, acting on DNA / interstrand cross-link repair / condensed chromosome / DNA polymerase binding / condensed nuclear chromosome / cellular response to ionizing radiation / male germ cell nucleus / meiotic cell cycle / cellular response to gamma radiation / protein-DNA complex / PML body / double-strand break repair via homologous recombination / HDR through Homologous Recombination (HRR) / Meiotic recombination / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / response to toxic substance / single-stranded DNA binding / site of double-strand break / double-stranded DNA binding / DNA recombination / chromosome, telomeric region / response to xenobiotic stimulus / mitochondrial matrix / DNA repair / DNA damage response / chromatin binding / centrosome / chromatin / nucleolus / perinuclear region of cytoplasm / enzyme binding / protein-containing complex / mitochondrion / nucleoplasm / ATP binding / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||||||||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) / Homo sapiens (human) / synthetic construct (others) /  | ||||||||||||||||||
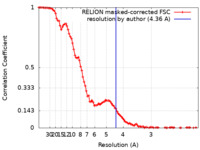
| Method | single particle reconstruction / cryo EM / Resolution: 4.36 Å | ||||||||||||||||||
 Authors Authors | Shioi T / Hatazawa S / Ogasawara M / Takizawa Y / Kurumizaka H | ||||||||||||||||||
| Funding support |  Japan, 5 items Japan, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2024 Journal: Nature / Year: 2024Title: Cryo-EM structures of RAD51 assembled on nucleosomes containing a DSB site. Authors: Takuro Shioi / Suguru Hatazawa / Eriko Oya / Noriko Hosoya / Wataru Kobayashi / Mitsuo Ogasawara / Takehiko Kobayashi / Yoshimasa Takizawa / Hitoshi Kurumizaka /  Abstract: RAD51 is the central eukaryotic recombinase required for meiotic recombination and mitotic repair of double-strand DNA breaks (DSBs). However, the mechanism by which RAD51 functions at DSB sites in ...RAD51 is the central eukaryotic recombinase required for meiotic recombination and mitotic repair of double-strand DNA breaks (DSBs). However, the mechanism by which RAD51 functions at DSB sites in chromatin has remained elusive. Here we report the cryo-electron microscopy structures of human RAD51-nucleosome complexes, in which RAD51 forms ring and filament conformations. In the ring forms, the N-terminal lobe domains (NLDs) of RAD51 protomers are aligned on the outside of the RAD51 ring, and directly bind to the nucleosomal DNA. The nucleosomal linker DNA that contains the DSB site is recognized by the L1 and L2 loops-active centres that face the central hole of the RAD51 ring. In the filament form, the nucleosomal DNA is peeled by the RAD51 filament extension, and the NLDs of RAD51 protomers proximal to the nucleosome bind to the remaining nucleosomal DNA and histones. Mutations that affect nucleosome-binding residues of the RAD51 NLD decrease nucleosome binding, but barely affect DNA binding in vitro. Consistently, yeast Rad51 mutants with the corresponding mutations are substantially defective in DNA repair in vivo. These results reveal an unexpected function of the RAD51 NLD, and explain the mechanism by which RAD51 associates with nucleosomes, recognizes DSBs and forms the active filament in chromatin. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38232.map.gz emd_38232.map.gz | 4.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38232-v30.xml emd-38232-v30.xml emd-38232.xml emd-38232.xml | 17.3 KB 17.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_38232_fsc.xml emd_38232_fsc.xml | 10 KB | Display |  FSC data file FSC data file |
| Images |  emd_38232.png emd_38232.png | 114.7 KB | ||
| Filedesc metadata |  emd-38232.cif.gz emd-38232.cif.gz | 5.9 KB | ||
| Others |  emd_38232_half_map_1.map.gz emd_38232_half_map_1.map.gz emd_38232_half_map_2.map.gz emd_38232_half_map_2.map.gz | 65.4 MB 65.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38232 http://ftp.pdbj.org/pub/emdb/structures/EMD-38232 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38232 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38232 | HTTPS FTP |
-Related structure data
| Related structure data |  8xbxMC  8jndC  8jneC  8jnfC  8xbtC  8xbuC  8xbvC  8xbwC  8xbyC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_38232.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38232.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_38232_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_38232_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : RAD51-nucleosome complex
| Entire | Name: RAD51-nucleosome complex |
|---|---|
| Components |
|
-Supramolecule #1: RAD51-nucleosome complex
| Supramolecule | Name: RAD51-nucleosome complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: DNA (5'-D(P*AP*AP*CP*GP*AP*AP*AP*AP*CP*GP*GP*CP*CP*AP*CP*CP*AP*CP...
| Macromolecule | Name: DNA (5'-D(P*AP*AP*CP*GP*AP*AP*AP*AP*CP*GP*GP*CP*CP*AP*CP*CP*AP*CP*G)-3') type: dna / ID: 1 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 48.595086 KDa |
| Sequence | String: (DA)(DT)(DC)(DA)(DG)(DA)(DA)(DT)(DC)(DC) (DC)(DG)(DG)(DT)(DG)(DC)(DC)(DG)(DA)(DG) (DG)(DC)(DC)(DG)(DC)(DT)(DC)(DA)(DA) (DT)(DT)(DG)(DG)(DT)(DC)(DG)(DT)(DA)(DG) (DA) (DC)(DA)(DG)(DC)(DT)(DC) ...String: (DA)(DT)(DC)(DA)(DG)(DA)(DA)(DT)(DC)(DC) (DC)(DG)(DG)(DT)(DG)(DC)(DC)(DG)(DA)(DG) (DG)(DC)(DC)(DG)(DC)(DT)(DC)(DA)(DA) (DT)(DT)(DG)(DG)(DT)(DC)(DG)(DT)(DA)(DG) (DA) (DC)(DA)(DG)(DC)(DT)(DC)(DT)(DA) (DG)(DC)(DA)(DC)(DC)(DG)(DC)(DT)(DT)(DA) (DA)(DA) (DC)(DG)(DC)(DA)(DC)(DG)(DT) (DA)(DC)(DG)(DC)(DG)(DC)(DT)(DG)(DT)(DC) (DC)(DC)(DC) (DC)(DG)(DC)(DG)(DT)(DT) (DT)(DT)(DA)(DA)(DC)(DC)(DG)(DC)(DC)(DA) (DA)(DG)(DG)(DG) (DG)(DA)(DT)(DT)(DA) (DC)(DA)(DC)(DC)(DC)(DA)(DA)(DG)(DA)(DC) (DA)(DC)(DC)(DA)(DG) (DG)(DC)(DA)(DC) (DG)(DA)(DG)(DA)(DC)(DA)(DG)(DA)(DA)(DA) (DA)(DA)(DA)(DA)(DC)(DA) (DA)(DC)(DG) (DA)(DA)(DA)(DA)(DC)(DG)(DG)(DC)(DC)(DA) (DC)(DC)(DA)(DC)(DG) |
-Macromolecule #2: DNA (5'-D(P*CP*GP*TP*GP*GP*TP*GP*GP*CP*CP*GP*TP*TP*TP*TP*CP*GP*TP...
| Macromolecule | Name: DNA (5'-D(P*CP*GP*TP*GP*GP*TP*GP*GP*CP*CP*GP*TP*TP*TP*TP*CP*GP*TP*T)-3') type: dna / ID: 2 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 48.952062 KDa |
| Sequence | String: (DC)(DG)(DT)(DG)(DG)(DT)(DG)(DG)(DC)(DC) (DG)(DT)(DT)(DT)(DT)(DC)(DG)(DT)(DT)(DG) (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DC)(DT) (DG)(DT)(DC)(DT)(DC)(DG)(DT)(DG)(DC)(DC) (DT) (DG)(DG)(DT)(DG)(DT)(DC) ...String: (DC)(DG)(DT)(DG)(DG)(DT)(DG)(DG)(DC)(DC) (DG)(DT)(DT)(DT)(DT)(DC)(DG)(DT)(DT)(DG) (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DC)(DT) (DG)(DT)(DC)(DT)(DC)(DG)(DT)(DG)(DC)(DC) (DT) (DG)(DG)(DT)(DG)(DT)(DC)(DT)(DT) (DG)(DG)(DG)(DT)(DG)(DT)(DA)(DA)(DT)(DC) (DC)(DC) (DC)(DT)(DT)(DG)(DG)(DC)(DG) (DG)(DT)(DT)(DA)(DA)(DA)(DA)(DC)(DG)(DC) (DG)(DG)(DG) (DG)(DG)(DA)(DC)(DA)(DG) (DC)(DG)(DC)(DG)(DT)(DA)(DC)(DG)(DT)(DG) (DC)(DG)(DT)(DT) (DT)(DA)(DA)(DG)(DC) (DG)(DG)(DT)(DG)(DC)(DT)(DA)(DG)(DA)(DG) (DC)(DT)(DG)(DT)(DC) (DT)(DA)(DC)(DG) (DA)(DC)(DC)(DA)(DA)(DT)(DT)(DG)(DA)(DG) (DC)(DG)(DG)(DC)(DC)(DT) (DC)(DG)(DG) (DC)(DA)(DC)(DC)(DG)(DG)(DG)(DA)(DT)(DT) (DC)(DT)(DG)(DA)(DT) |
-Macromolecule #3: DNA repair protein RAD51 homolog 1
| Macromolecule | Name: DNA repair protein RAD51 homolog 1 / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.291398 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSHMAMQMQL EANADTSVEE ESFGPQPISR LEQCGINAND VKKLEEAGFH TVEAVAYAPK KELINIKGIS EAKADKILAE AAKLVPMGF TTATEFHQRR SEIIQITTGS KELDKLLQGG IETGSITEMF GEFRTGKTQI CHTLAVTCQL PIDRGGGEGK A MYIDTEGT ...String: GSHMAMQMQL EANADTSVEE ESFGPQPISR LEQCGINAND VKKLEEAGFH TVEAVAYAPK KELINIKGIS EAKADKILAE AAKLVPMGF TTATEFHQRR SEIIQITTGS KELDKLLQGG IETGSITEMF GEFRTGKTQI CHTLAVTCQL PIDRGGGEGK A MYIDTEGT FRPERLLAVA ERYGLSGSDV LDNVAYARAF NTDHQTQLLY QASAMMVESR YALLIVDSAT ALYRTDYSGR GE LSARQMH LARFLRMLLR LADEFGVAVV ITNQVVAQVD GAAMFAADPK KPIGGNIIAH ASTTRLYLRK GRGETRICKI YDS PCLPEA EAMFAINADG VGDAKD UniProtKB: DNA repair protein RAD51 homolog 1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)